Journal:Screening for more than 1,000 pesticides and environmental contaminants in cannabis by GC/Q-TOF
| Full article title | Screening for more than 1,000 pesticides and environmental contaminants in cannabis by GC/Q-TOF |
|---|---|
| Journal | Medical Cannabis and Cannabinoids |
| Author(s) | Wylie, P.L.; Westland, J.; Wang, M.; Radwan, M.M.; Majumdat, C.G.; ElSohly, M.A. |
| Author affiliation(s) | Agilent Technologies, University of Mississippi, ElSohly Laboratories |
| Primary contact | Email: Philip dot l dot wylie at gmail dot com |
| Year published | 2020 |
| Volume and issue | 3(1) |
| Page(s) | 14–24 |
| DOI | 10.1159/000504391 |
| ISSN | 2504-3889 |
| Distribution license | Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International |
| Website | https://www.karger.com/Article/FullText/504391 |
| Download | https://www.karger.com/Article/Pdf/504391 (PDF) |
Abstract
A method has been developed to screen cannabis extracts for more than 1,000 pesticides and environmental pollutants using gas chromatography coupled to a high-resolution accurate mass quadrupole time-of-flight mass spectrometer (GC/Q-TOF). An extraction procedure was developed using acetonitrile with solid-phase extraction cleanup. Before analysis, extracts were diluted 125:1 with solvent. Two data mining approaches were used together with a retention-time-locked Personal Compound Database and Library (PCDL) containing high-resolution accurate mass spectra for pesticides and other environmental pollutants. In the first approach, a Find-by-Fragments (FbF) software tool extracted several characteristic exact mass ions within a small retention time window where the compound eluted. For each compound in the PCDL, the software evaluated the peak shape and retention time of each ion, as well as the monoisotopic exact mass, ion ratios, and other factors to decide if the compound was present or not. In the second approach, Unknowns Analysis (UA) software with a peak-finding algorithm called SureMass was used to deconvolute peaks in the chromatogram. The accurate mass spectra were searched against the PCDL using spectral matching and retention time as filters. A subset PCDL was generated containing only pesticides that are most likely to be found on foods in the US. With about 250 compounds in the smaller PCDL, there were fewer hits for non-pesticides, and data review was much faster. Organically grown cannabis was used for method development. Twenty-one confiscated cannabis samples were analyzed and ten were found to have no detectable pesticides. The remaining 11 samples had at least one pesticide, and one sample had seven detectable residues. Quantitative analysis was run on the confiscated samples for a subset of the pesticides found by screening. Two cannabis samples had residues of carbaryl and malathion that were estimated to be about 10 times greater than the highest U.S. Environmental Protection Agency (EPA) tolerance set for food and about 4,000 times greater than the Canadian maximum residue limits for dried cannabis flower.
Introduction
The use of medicinal cannabis is legal in 20 countries and 33 U.S. states, while two countries, Canada and Uruguay, have fully legalized the sale and use of recreational cannabis nationwide.[1] In the U.S., 11 states and the District of Columbia have legalized recreational use of cannabis, but it is still illegal at the federal level. Many countries and U.S. states have decriminalized possession and use of cannabis.[2] With so many people having access to cannabis, there is an increased need to test cannabis products to ensure their safety. Cannabis plants are subject to various pests and diseases which may require the judicious use of pesticides to maintain plant health. McPartland has published two comprehensive reviews on the diseases[3] and pests[4] that attack Cannabis plants. But, pesticide residues on the plant material are particularly concerning because cannabis can be ingested, smoked, or extracted and concentrated for use in everything from food and beverages to tinctures and suppositories. Many jurisdictions that have legalized cannabis require testing for pesticide residues. For example, Oregon[5], California[6], and Canada[7] have lists of 59, 66, and 95 pesticides, respectively, that must be targeted by analysts. However, some growers use pesticides that are not on lists of acceptable compounds for which maximum residue limits (MRLs) have been set. Product recalls are common, companies have gone out of business, and occasionally, someone is fined for the misuse of pesticides. On top of this, the unregulated market for cannabis is still much bigger than the legal market.[8] Many of the illegal growers use pesticides carelessly and leave unknown levels of sometimes illegal pesticides on the plant material. One real concern is the affect that pesticides and cannabinoids might have on a developing fetus.[9]
Typically, laboratories test for pesticide residues on cannabis using gas chromatography and liquid chromatography with tandem quadrupole detectors (GC/MS/MS and LC/MS/MS). These instruments are extremely sensitive and are very selective in the multiple reaction monitoring mode. However, they can only find those pesticides that are on the target list. Other pesticides and environmental contaminants will be missed.
Clandestine cannabis growers often use illegal pesticides at their grow sites. For example, carbofuran, a carbamate insecticide that is now banned for use in the U.S., was found at 78% of eradicated illegal grow sites in 2017.[10][11] This highly toxic pesticide would be missed by typical laboratory testing procedures employing GC/MS/MS and LC/MS/MS for target compound analysis.
Because cannabis has been illegal in most countries around the world, there are not many studies available that evaluate cannabis samples for pesticide residues. A recent paper describes three sample preparation methods for cannabis leaves, dried flowers, and oils with analysis of pesticide residues by HPLC/MS/MS, GC/MS/MS, and GC/MS. One hundred and forty-four samples of cannabis leaves, dried flowers, and oils were obtained from Canadian dispensaries and were tested using their validated methods. Of 26 samples that contained unauthorized pesticides, myclobutanil was found most often (20 samples) followed by the acaricide bifenazate in nine samples.[12] Schneider et al.[13] analyzed 50 samples of confiscated cannabis for 160 pesticides by UHPLC/MS/MS and GC/MS in the scan mode. Seven different pesticides were found in 19 samples. A headspace solid-phase microextraction GC/MS method has been tested for the quantitative analysis of nine pesticides in cannabis.[14]
Here, we describe a procedure for suspect screening using gas chromatography coupled to a high-resolution accurate mass quadrupole time-of-flight mass spectrometer (GC/Q-TOF) together with a Pesticides and Environmental Pollutants (P&EP) Personal Compound Database and Library (PCDL). The PCDL contains chemical formulas, isotope patterns, and electron ionization mass spectra with accurate monoisotopic mass assignments for more than 1,000 GC-amenable pesticides and environmental contaminants. The method allows one to presumptively identify contaminants without the need to purchase analytical reference standards. Of course, standards are required for positive identification or for quantitative analysis. The procedure is qualitative in nature, but quantification is possible when standards are available. The GC/Q-TOF, together with a pesticide PCDL, have been used for the detection of pesticides in aquatic environments[15] and in various foods[16][17], but it has never been used to analyze pesticide residues in cannabis. While this paper describes the analysis of GC-amenable pesticide residues on cannabis, there is a need for a similar broad screening method using high-resolution accurate mass liquid chromatography–mass spectrometry (LC–MS) because most (but not all) pesticides are LC-amenable.
This method is intended to help analysts such as government regulators, researchers, and other labs find pesticides on cannabis samples that may not be on a laboratory’s normal target list. As new cannabis companies establish their brand name, and as established companies move into the cannabis market, there will be greater incentive to protect their brand reputation by making sure that no unapproved pesticides contaminate their products.
Materials and methods
Source of cannabis
Samples of ground cannabis flower were obtained from the University of Mississippi Marijuana Project, which has grown all the marijuana for U.S. Government-approved research. The University of Mississippi Marijuana Project also has access to thousands of confiscated cannabis samples, which are used for potency assessments.[18][19][20] Two kinds of cannabis samples were studied: (1) ground cannabis flower from cannabis grown at the University of Mississippi using organic farming practices so no pesticides were used in production, and (2) ground samples of confiscated cannabis for which the origin and growing conditions are unknown. The organically grown cannabis samples were used for method development, while the confiscated samples were used to assess the extent of pesticide contamination of illicit cannabis. All the cannabis samples were stored in the dark at room temperature for months or years, so labile pesticides that may have been on the samples are likely to have decomposed.
Solvents and standards
ACS/HPLC grade acetonitrile was purchased from Burdick & Jackson (Muskegon, MI, USA). HPLC grade hexane was purchased from Sigma Aldrich (St. Louis, MO, USA), and environmental grade acetone was purchased from Alfa Aesar (Ward Hill, MA, USA). Pesticide standards were purchased from Agilent Technologies, Inc. (Santa Clara, CA, USA).
Sample preparation
Cannabis is an extremely complex plant containing a large variety of chemical compounds, including terpenes, carbohydrates, fatty acids and their esters, amides, amines, phytosterols, phenolic compounds, and cannabinoids.[21] The complexity of the cannabis matrix makes detection and accurate quantification of trace levels of pesticides more challenging. Interfering compounds can negatively impact ionization in the mass spectrometer, affect signal-to-noise ratios, and build-up in the instrument source and GC column, thus decreasing productivity and increasing maintenance. To overcome this challenge, a combination of optimized sample preparation and state-of-the-art instrumentation is required.
Samples were prepared using the method described for GC/MS/MS analysis in a recently published paper by Roy et al.[22] This procedure uses a pass through SPE cleanup (SampliQ C18 Endcapped SPE Cartridge, Agilent Technologies) followed by a 125:1 dilution. The objective of the cleanup is to remove as much matrix as possible without removing or destroying any pesticides in the sample. Recoveries and %RSD values have been published for the 95 pesticides for which Health Canada requires testing. Recoveries for all but three compounds fell in the 70–110% range, and all RSD values were less than 6%.[22]
Preparation of matrix-matched calibration standards
Post-extraction calibration standards were prepared in pesticide-free cannabis extract (1 g in 25 mL acetonitrile), which was further diluted 5:1 with 50:50 (v/v) hexane:acetone (acidified with 0.1% formic acid). This resulted in dilution of the cannabis matrix by a factor of 125:1. Matrix-matched pesticide calibration standards were prepared at 50, 25, 10, 5, 2.5, 1, 0.8, 0.5, and 0.3 ng/g from a 1,000 ng/g stock solution containing either the California pesticide list[4] or the Oregon list.[3]
Instrumentation
Samples were analyzed on an Agilent 8890 GC equipped with an Agilent 7693 Autosampler, a multimode inlet (MMI), a pneumatic switching device, a Purged Ultimate Union (PUU) for backflushing[23][24], and two 15 m × 0.25 mm × 0.25 µm Agilent HP-5MS Ultra Inert columns. The mass spectral detector was an Agilent 7250 High Resolution Accurate Mass Q-TOF MS operated in the full spectral acquisition mode.
Column 1 was connected between the MMI and the PUU, while column 2 was connected between the PUU and the mass spectrometer transfer line. The helium carrier gas was set to the constant flow mode. The nominal column 1 flow rate was 1.2 mL/min, and the flow rate in column 2 was set at 0.2 mL/min higher than column 1 (nominally 1.4 mL/min). The exact flow rates were adjusted to lock the retention time of chlorpyrifos-methyl to 9.143 min. The inlet was held at 280°C and was operated in the pulsed splitless mode with a pulse pressure of 25 psi lasting for 0.5 min. The purge vent was opened at 0.7 min with a flow of 50 mL/min until 2 min, when it was switched to 20 mL/min using the gas saver function. The septum purge flow was 3 mL/min and was set to the switched mode. An Agilent Technologies 4 mm i.d. single tapered Ultra Inert liner with deactivated glass wool (PN 5190-2293) was installed in the inlet. The GC oven temperature was held at 60°C for 1 min, programmed at 40°C/min to 170°C, held for 0 min, then ramped at 10°C/min to 310°C and held for 3 min. At the end of the run with the oven at 310°C, column 1 was backflushed for 2 min at –7.874 mL/min by reducing the inlet pressure to 2 psi and increasing the pneumatic switching device pressure to 60 psi.
The Q-TOF mass spectrometer was equipped with a high-emission low-energy-capable electron ionization source, which was operated with an electron energy of 70 eV and an emission current of 5.0 µA. The source, quadrupole, and transfer line temperatures were set to 280°C, 150°C, and 300°C, respectively. Data were acquired at a rate of 5 Hz from 45 to 550 m/z with a 3 min solvent delay. Collision cell flows were set to 4.0 mL/min and 1.0 mL/min for helium and nitrogen, respectively. Automated TOF mass calibration was performed after every second injection using a keyword command in the sequence table. Data were acquired using Agilent MassHunter Acquisition Software (B.10.0). Data analysis was performed using Agilent MassHunter Qualitative (B.10.0) and Quantitative (B.10.0) analysis software.
Results and discussion
Complexity of the cannabis matrix
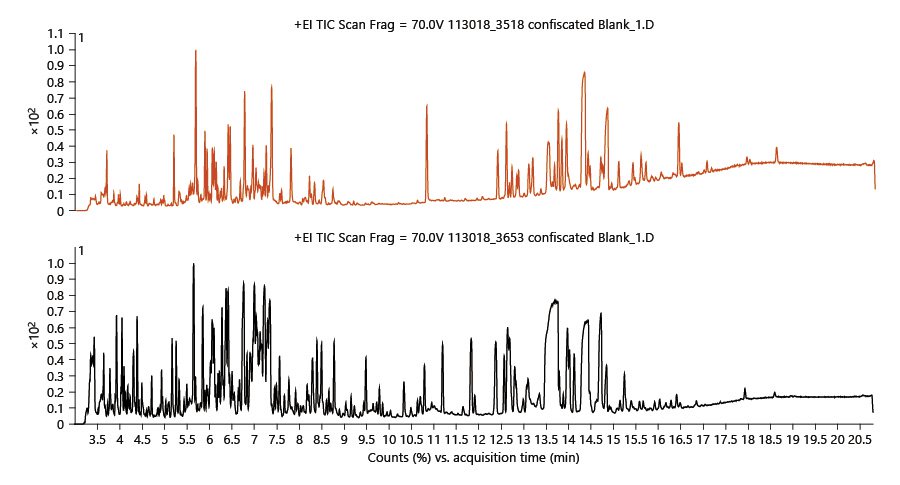
Over 100 different biosynthesized cannabinoids have been identified in cannabis, which constitute 10–30% of the plant. Typical cannabis flowers for recreational use are reported to contain a mean concentration of Δ9-tetrahydrocannabinol (Δ9-THC) or its precursor tetrahydrocannabinolic acid (THCA) of 17.1%.[18] More than 560 constituents, from diverse chemical classes, have been identified in cannabis.[21][25] Many of these compounds are extracted using typical pesticide extraction methods, which are only partially successful in removing these endogenous phytochemicals. Cannabis is a very complex matrix that requires a tailored sample preparation method to help separate the matrix from the pesticides. After the SPE passthrough cleanup and dilution, there is still a need for instrumentation with high sensitivity and selectivity to accurately identify and quantify these trace pesticides. Figure 1 shows two typical GC/Q-TOF chromatograms of cannabis extracts showing how complex the samples are even after SPE cleanup and dilution. To maintain a clean source and remove non-volatile compounds, column 1 was backflushed at the end of the run for 2 min at –7.874 mL/min.
|
Pesticides and environmental pollutants PCDL
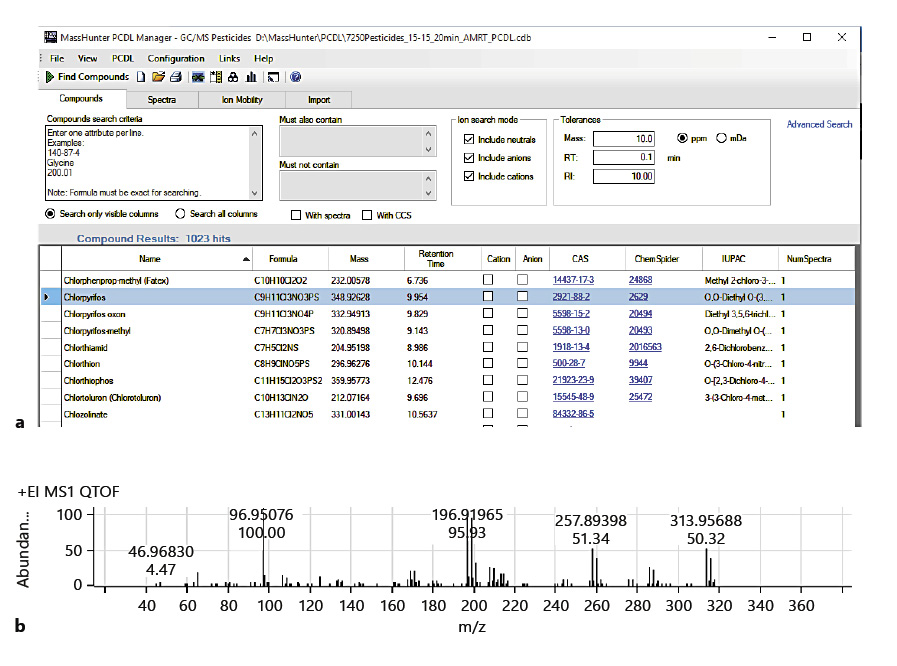
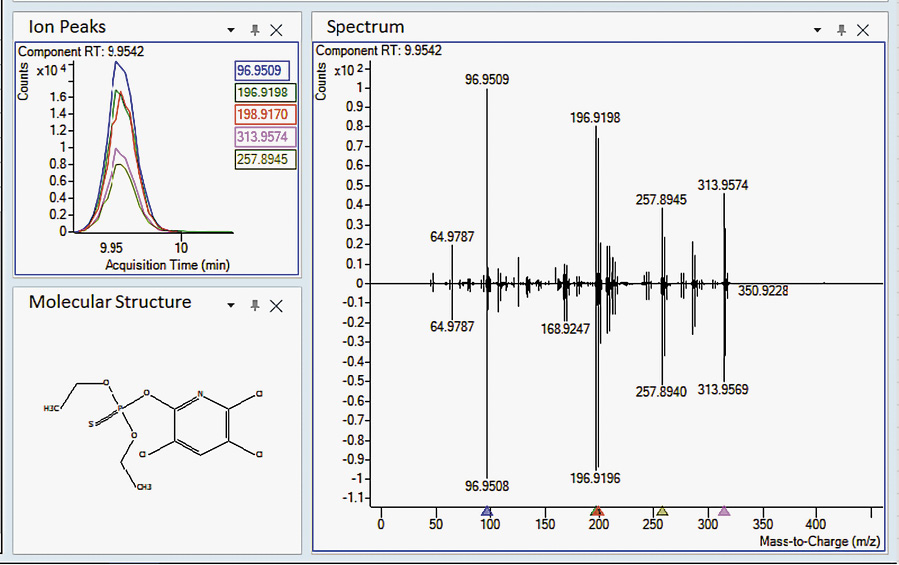
A commercially available PCDL (Agilent Technologies) was used in two different data analysis approaches: Find-by-Fragments (FbF) and Unknowns Analysis (UA). Most entries in this PCDL are pesticides, but other environmental pollutants (e.g., pesticide metabolites, fire retardants, PAHs, phthalates, chlorobenzenes, and nitrosamines) are included. Each entry contains the compound’s locked retention time[26] and a curated accurate mass spectrum, with the accurate mass assigned for each spectral peak. Figure 2a shows a section of the table, and Figure 2b shows the spectrum of chlorpyrifos, one of the pesticide entries. Though not shown in Figure 2, the molecular structure is included for each entry.
|
Suspect screening for all of the PCDL compounds
The FbF screening tool in the qualitative analysis software was used to screen data files for all 1,020 compounds included in the P&EP PCDL. The software looks at the first entry in the PCDL and chooses several of the most significant ions in the spectrum (based on molecular weight and abundance). It then extracts those ions over a small range around the compound’s locked retention time (which it also reads from the PCDL). In theory, ions belonging to the same molecule should have identical retention times and the same relative peak shape. The software chooses one ion and compares the other [[|Mass chromatography#Extracted-ion chromatogram (XIC)|extracted-ion chromatograms]] (EICs) to this “reference ion.” Finally, it scores each EIC based on the similarity of peak shape and retention time to the reference ion. An EIC with a “coelution score” greater than a value set by the analyst is said to be “qualified.” Before running FbF, the user chooses the PCDL to be used, and certain other parameters such as the mass accuracy window for the EICs, the number of ions to extract, the size of the retention time window, the nature of the charge carrier, the number of qualified ions required to call the compound a “hit,” and a few other parameters as shown in Table S1 (see Supplementary material). The software does this for all entries in the PCDL and creates a summary report for the compounds it found.
Reducing false positive or insignificant hits using a subset PCDL
Under certain circumstances, the FbF algorithm can produce numerous “hits,” some of which may be false positives or not useful to the analyst. The full P&EP PCDL contains many non-pesticide compounds that are classified as environmental pollutants such as phthalates, PAHs, chlorobenzenes, synthetic musk compounds, and fire retardants. These often show up in samples but may not be useful to the analyst only looking for pesticide residues. Also, many of the pesticides in the PCDL are not commonly found on fruits and vegetables, and one might assume that they would be less likely to be found on cannabis. The U.S. Food and Drug Administration's (FDA) Pacific Northwest Regional Laboratory (PNRL) in Bothell, WA, screens food commodities for pesticide residues by GC/TQ. For this method, they only screen for pesticides that have been seen in the past or that were recently approved by the U.S. Environmental Protection Agency (EPA).[27] They screen for a much larger set of pesticides using a single quadrupole GC/MS with deconvolution and a retention-time-locked database.[28] Any new compounds they find are then added to the GC/TQ method. They have a list of pesticides that they run on their GC/TQ that are most likely to be encountered when screening both imported and domestic foods. Drawing from the PNRL list—together with the California, Oregon, and Canada target lists—a subset PCDL was made that contains about 250 pesticides. This allowed for a broad screen of likely pesticides without the environmental pollutants and rarely encountered pesticides. Samples were analyzed using both the complete PCDL and the smaller subset PCDL. The subset PCDL reduced the number of hits needing to be reviewed from an average of 71 to 15 per sample (see Table S2, Supplementary material).
Interpreting the Find-by-Fragments results
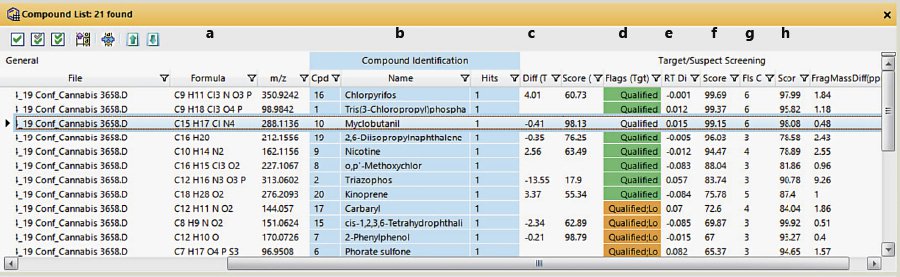
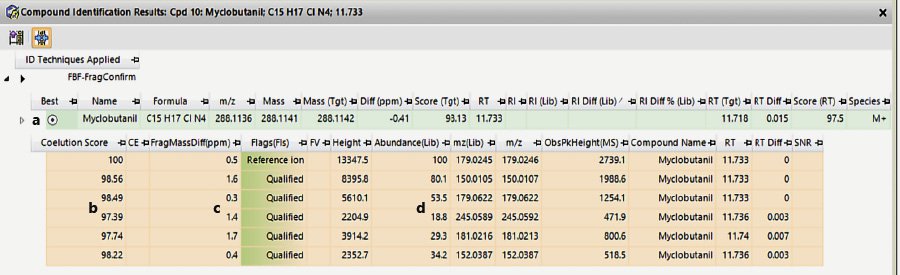
The FbF algorithm returns several tables and plots (Fig. 3-5) that provide evidence for compounds that it has identified as probable hits. Figures 3-5 show results from analyzing a cannabis extract (diluted 125:1) that was spiked at 5 ppb with 66 pesticides (not all of which are GC-able). The table in Figure 3 lists all the compounds meeting the user-defined qualifying requirements for a hit. This table provides information about the compound and the confidence in its identification. In addition to the name of the compound, it gives the measured retention time, the measured monoisotopic molecular weight (if a molecular ion is present), and the difference between the measured and theoretical MW (Fig. 3, column c). The difference between the PCDL retention time and the measured value is shown in column e, and, because of retention time locking[26], this value is almost always <0.1 min. Column g indicates how many of the six extracted ions were qualified for each compound listed. Myclobutanil and chlorpyrifos are two pesticides for which all six extracted ions were qualified, indicating a confident match. Some pesticides only have a few distinctive and abundant ions in the spectrum. When the concentration gets low, some of the less abundant ions will not be found, and interferences from endogenous compounds become more important. This is especially true with cannabis extracts because they are so dirty.
|
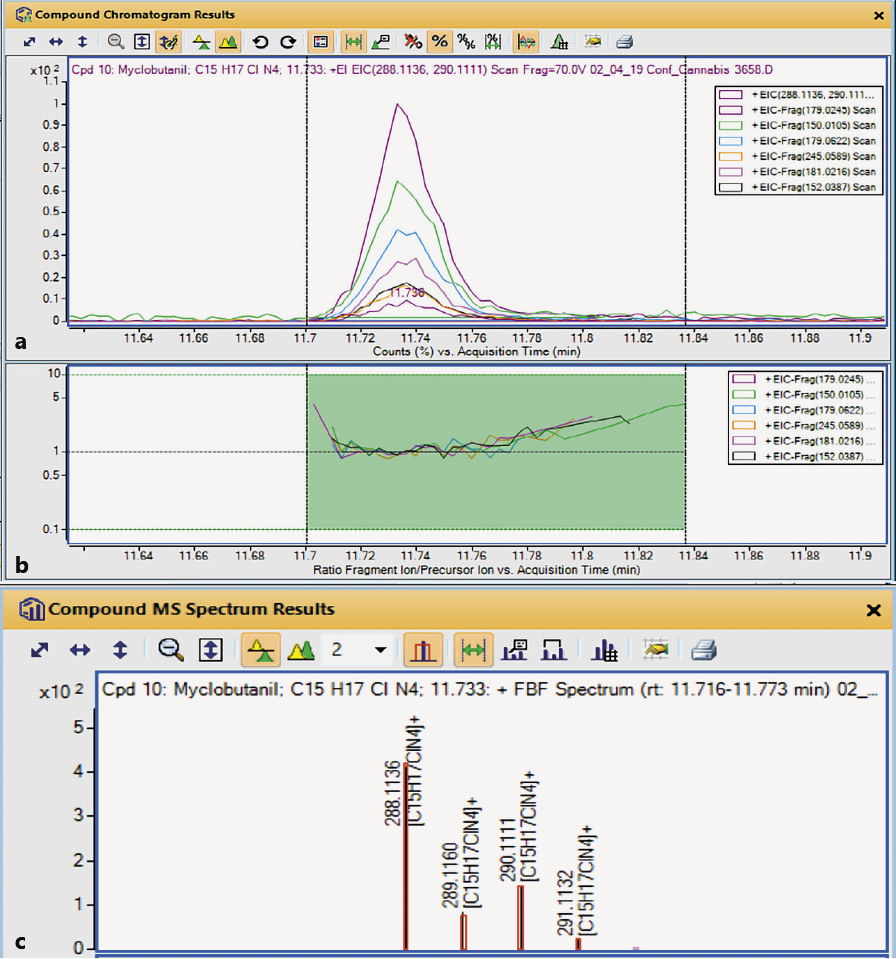
Figure 4a shows accurate mass EIC for the characteristic ions chosen by the software. If all the ions come from the same compound, and there are no interferences, the RT and relative abundance of each ion across the full peak width should be identical. Figure 4b shows a coelution plot in which the relative abundance of each EIC is normalized to the response of the reference peak. The plot in Figure 4b would be a horizontal straight line if the normalized relative abundances were identical across the peak. When the molecular ion is present in measurable abundance, the software shows a plot of the M+· isotope pattern (Fig. 4c). Isotope spacing (difference in m/z) and relative abundance are plotted (black lines) and compared to the theoretical values (red boxes). When the spacing and relative abundances are a good match for the theoretical values, this provides strong evidence that a compound with this molecular formula is the correct choice.
|
Figure 5 gives more information about the individual extracted ions and provides a summary of the compound information in the top line (highlighted in light green). This is another place to see the closeness of the measured MW and RT to the database values. For each extracted ion it provides a coelution score for which the minimum acceptable score in this method was 70. For each ion, its measured m/z value is compared to values taken from the database spectrum and the difference is shown in ppm (Figure 5c).
|
Pesticide detection limits
While this GC/Q-TOF method would most likely be used as a qualitative method to detect pesticides that have not been approved for use on cannabis, it is useful to know the approximate detection limits that can be achieved for various pesticides. Calibration samples were prepared at nine levels, from 0.3 to 50 ppb, in pesticide-free cannabis matrix that was diluted 125:1. The samples were spiked with 66 pesticides regulated for cannabis by California laws.[6] These were analyzed using the FbF approach to see how well the algorithm works as the pesticide concentrations decrease. The FbF algorithm was run on samples at each spiking level, with six ions extracted for each compound and two qualified ions required for a hit. At the highest spiking level, 50 ppb in the diluted extract, the software found all 45 of the GC-amenable compounds (see Table S3, Supplementary material). At 5 ppb, the software was still able to find 40 of them. When the concentration in the diluted extract is 0.8 ppb, this is equivalent to 100 ppb in the original cannabis sample, assuming 100% recovery during extraction. This corresponds to the lowest detection limit required by California; however, this is not low enough to meet many of the Canadian tolerances for cannabis.[7] At this level, 28 (62%) of the compounds could be tentatively identified. Twenty-one of the pesticides could still be identified at the lowest spiking level tested (0.3 ppb).
As the pesticide concentration gets lower, the number of qualified ions tends to fall off. Less abundant ions in the spectrum often cannot be pulled out of the high background noise from the cannabis matrix. In most cases, this is not for a lack of instrument resolution. It is because an interference has the identical mass as the target ion. At the 50-ppb spiking level, all six ions were found for 30 pesticides. At 5 ppb, 13 compounds had all six ions qualified. Only bifenthrin and dichlorvos still had six qualified ions at the lowest spiking level. Table S3 (see Supplementary material) shows the number of ions found for those pesticides that could be detected at each calibration level.
Pesticide identification using UA
MassHunter UA software uses an algorithm called “SureMass” to find peaks in the accurate mass chromatogram and searches a mass spectral library or PCDL to identify the peaks. If the library has locked retention times or retention index values, these can be used as filters. In this case, hits must have the correct RT and be a good match to the database spectrum. The SureMass peak-finding algorithm takes advantage of the added information available in high-resolution accurate mass data. Peaks that were identified by SureMass were searched against the same PCDLs used for the FbF approach. Figure 6 shows results for the identification of chlorpyrifos in one of the confiscated cannabis samples. This approach is inherently simpler than the FbF method, and data review is relatively easy. As shown in Table S2 (see Supplementary material), UA produces many fewer hits to be reviewed when using either PCDL. In most cases, after review, FbF and UA gave the same results.
|
Analysis of confiscated cannabis samples
Cannabis that is confiscated by authorities is most likely purchased through the unregulated market or grown by the user, so one does not know which, if any, pesticides were used on the plants. Many reports indicate that illegal growers use pesticides indiscriminately and often use pesticides, such as carbofuran, that are banned in the U.S.[10][11] The University of Mississippi Marijuana Project has access to thousands of confiscated cannabis samples. We chose 21 different samples of confiscated cannabis flower for pesticide suspect screening by GC/Q-TOF.
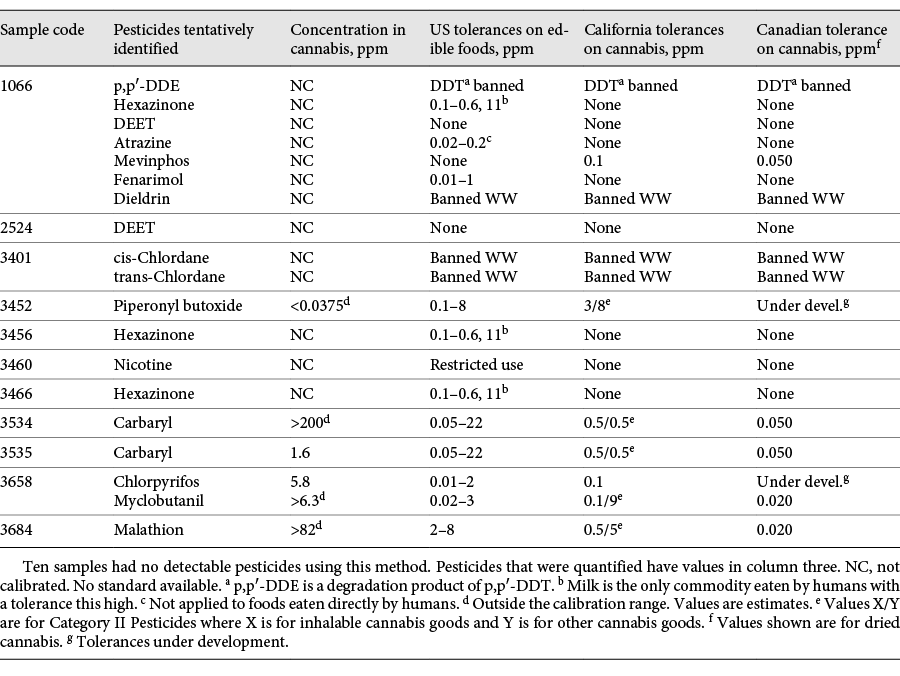
Of the 21 confiscated cannabis samples analyzed, 10 were found to have no detectable pesticides using both FbF and UA. Two samples were found to have two pesticides each, and eight samples had a single pesticide. One sample (#1066) had seven different pesticide, herbicide, and fungicide residues: p,p′-DDE, hexazinone, DEET, atrazine, mevinphos, fenarimol, and dieldrin (Table 1).
|
Table 1 shows the pesticides, herbicides, and fungicides that were tentatively identified in eleven of the of the 21 confiscated cannabis samples. Some of the halogenated pesticides in the table such as p,p′-DDE (a degradation product of p,p′-DDT), dieldrin, and chlordane have been banned in the U.S. for many years. It is likely that the cannabis samples having these contaminants were grown in soil that was previously contaminated by these persistent pollutants.
Calibration curves were prepared from 0.3 to 50 ppb in pesticide-free cannabis extract (diluted 125:1) for some of the pesticides, and quantitative analysis was performed using the GC/Q-TOF. The concentrations shown in Table 1 are for the residues in the original cannabis sample, obtained by multiplying the measured amounts by the dilution factor (125). Only two samples had residue amounts that fell on the calibration curve – carbaryl in sample 3535 and chlorpyrifos in sample 3658. The others were either above or below the calibration levels used, and the values shown are only estimates. Ordinarily, one would dilute samples and reanalyze them when the observed concentration levels are above the calibration curve. That was not done in this case since the objective was to get an approximate idea of the contamination levels. One sample, 3534, was estimated to have well over 200 ppm of carbaryl, while another, 3684, showed high levels of malathion. For comparison, U.S. tolerance ranges are shown for these pesticides on various foods. Cannabis MRLs are also shown for California, which classifies pesticides into two categories. Category I pesticides should not be found on cannabis at all, and laboratories must demonstrate a limit of quantification of 0.10 µg/g, making this the de facto MRL. Category II pesticides may be used on cannabis, and there are two sets of MRLs for these: one for products that are intended to be inhaled and another for all other products. Canadian MRLs are generally lower than those in California. In all but one case, the measured residue levels in the confiscated cannabis exceeded the California and Canadian limits. Carbaryl in sample 3534 and malathion in sample 3684 were estimated to exceed the Canadian MRLs by a factor of 4,000.
Conclusions
A high-resolution, accurate mass GC/Q-TOF instrument was used to screen cannabis samples for more than 1,000 pesticides and other environmental contaminants. Each sample of ground dried cannabis flower was extracted in acetonitrile, and the extract was passed through a SampliQ C18 endcapped SPE cartridge. Because cannabis extracts are so dirty, they were diluted 125:1 with solvent. The GC method used was retention time-locked to a commercially available PCDL that contains accurate mass spectra and locked retention times for 1,020 compounds. To make data review easier, the original PCDL was used to create a subset PCDL containing about 250 pesticides that are most commonly found on food commodities in the U.S. Data files were reviewed using two different procedures for finding suspect compounds – FbF and UA. Twenty-one samples of confiscated cannabis flower were analyzed and eleven were found to be contaminated with detectable pesticide residues. Thirteen different contaminates were tentatively identified using these procedures. Concentrations of some of those contaminants were determined by calibration with standards. Two cannabis samples had pesticide levels that were estimated to be about 10 times greater than the highest EPA tolerance set for food in the U.S. and about 4,000 times greater than the Canadian MRLs for dried cannabis flower. The two approaches to finding pesticides—FbF and UA—gave very similar results when analyzing the confiscated cannabis samples, but data review was much easier using UA.
Supplementary material
Supplementary data is available for viewing and download at https://doi.org/10.6084/m9.figshare.11558607.v1. It lists Tables S1–S3 in a .docx file.
Acknowledgements
Author contributions
M.M. Radwan, C.G. Majumdar, and J. Westland developed the extraction procedure. M. Wang and P.L. Wylie worked on extractions and analyzed samples on the GC/Q-TOF. M.A. ElSohly helped with experimental design and manuscript review.
Funding
This research was partially supported by the National Institute on Drug Abuse (N01DA-15-7793.
Conflicts of interest
The authors have no financial or non-financial competing interest. P.L. Wylie and J. Westland are employed by Agilent Technologies, which provided the instrumentation and software used. Agilent products and solutions are intended to be used for cannabis quality control and safety testing in laboratories where such use is permitted under state/country law.
References
- ↑ Hassan, A. (17 October 2018). "All the places in the world you can (legally) smoke weed". Quartz. https://qz.com/1427177/where-is-marijuana-legal-around-the-world/. Retrieved 01 May 2019.
- ↑ "Map of Marijuana Legality by State". DISA Global Solutions. 2019. https://disa.com/map-of-marijuana-legality-by-state. Retrieved 01 May 2019.
- ↑ 3.0 3.1 McPartland, J.M. (1996). "A review of Cannabis diseases". Journal of the International Hemp Association 3 (1): 19–23. http://www.internationalhempassociation.org/jiha/iha03111.html.
- ↑ 4.0 4.1 McPartland, J.M. (1996). "Cannabis pests". Journal of the International Hemp Association 3 (2): 49, 52–55. http://www.internationalhempassociation.org/jiha/iha03201.html.
- ↑ Farrer, D.G. (December 2015). "Technical report: Oregon Health Authority’s process to decide which types of contaminants to test for in cannabis products, and levels for action" (PDF). Oregon Health Authority. https://www.oregon.gov/oha/ph/PreventionWellness/marijuana/Documents/oha-8964-technical-report-marijuana-contaminant-testing.pdf. Retrieved 01 May 2019.
- ↑ 6.0 6.1 "16 CCR § 5719 Residual Pesticides Testing". Westlaw. Thomson Reuters. https://govt.westlaw.com/calregs/Document/I8CCCCCCFBE19419D9DDFFA11E5E29042?viewType=FullText&originationContext=documenttoc&transitionType=CategoryPageItem&contextData=(sc.Default)#co_anchor_IB78AA9DD7002490FB0286370F626BEB. Retrieved 01 May 2019.
- ↑ 7.0 7.1 Health Canada (2019). "Mandatory cannabis testing for pesticide active ingredients - List and limits". Government of Canada. https://www.canada.ca/en/public-health/services/publications/drugs-health-products/cannabis-testing-pesticide-list-limits.html. Retrieved 01 May 2019.
- ↑ McGovern, S. (17 December 2018). "11 Facts Cannabis Entrepreneurs Should Know About the Black Market". Green Entrepreneur. https://www.greenentrepreneur.com/article/324679. Retrieved 01 May 2019.
- ↑ Leung, M.C.K.; Silva, M.H.; Palumbo, A.J. et al. (2019). "Adverse outcome pathway of developmental neurotoxicity resulting from prenatal exposures to cannabis contaminated with organophosphate pesticide residues". Reproductive Toxicology 85: 12–18. doi:10.1016/j.reprotox.2019.01.004. PMID 30668982.
- ↑ 10.0 10.1 Fimrite, P. (29 May 2018). "Illegal pot grows spread deadly pesticides, other hazards, despite change in law". San Francisco Chronicle. https://www.sfchronicle.com/green/article/Illegal-pot-grows-spread-deadly-pesticides-other-12952302.php. Retrieved 01 May 2020.
- ↑ 11.0 11.1 Thompson, C.M.; Gabriel, M.W.; Purcell, K.L. (2017). "An ever-changing ecological battlefield: marijuana cultivation and toxicant use in western forests". The Wildlife Professional 11 (3): 42–6. https://www.fs.usda.gov/treesearch/pubs/55041.
- ↑ Moulins, J.R.; Blais, M.; Montsion, K. et al. (2018). "Multiresidue Method of Analysis of Pesticides in Medical Cannabis". Journal of AOAC International 101 (6): 1948–60. doi:10.5740/jaoacint.17-0495. PMID 29843862.
- ↑ Schneider, S.; Bebing, R.; Dauberschmidt, C. (2013). "Detection of pesticides in seized illegal cannabis plants". Analytical Methods 6 (2): 515–20. doi:10.1039/C3AY40930A.
- ↑ Ilias, Y.; Rudaz, S.; Christen, P. et al. (2006). "Headspace Solid-Phase Microextraction of Pesticide Residues in Cannabis Samples". CHIMIA International Journal for Chemistry 60 (12): 846–51. doi:10.2533/chimia.2006.846.
- ↑ Moschet, C.; Lew, B.M.; Hasenbein, S. et al. (2017). "LC- and GC-QTOF-MS as Complementary Tools for a Comprehensive Micropollutant Analysis in Aquatic Systems". Environmental Science and Technology 51 (3): 1553-1561. doi:10.1021/acs.est.6b05352. PMC PMC7238889. PMID 328026950. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=PMC7238889.
- ↑ Chen, K.; Sanderson, J. (16 June 2017). "Screening of Pesticides and Other Contaminants in Food Matrices Using a Novel High‑resolution GC/Q-TOF with a Low‑energy‑capable EI Source" (PDF). Agilent Technologies, Inc. https://gcms.cz/labrulez-bucket-strapi-h3hsga3/paper/5991-8170EN.pdf.
- ↑ Li, J.-X.; Li, X.-Y.; Chang, Q.-Y. et al. (2018). "Screening of 439 Pesticide Residues in Fruits and Vegetables by Gas Chromatography-Quadrupole-Time-of-Flight Mass Spectrometry Based on TOF Accurate Mass Database and Q-TOF Spectrum Library". Journal of AOAC International 101 (5): 1631–8. doi:10.5740/jaoacint.17-0105. PMID 29724258.
- ↑ 18.0 18.1 Chandra, S.; Radwan, M.M.; Majumdar, C.G. et al. (2019). "New trends in cannabis potency in USA and Europe during the last decade (2008-2017)". European Archives of Psychiatry and Clinical Neuroscience 269 (1): 5–15. doi:10.1007/s00406-019-00983-5. PMID 30671616.
- ↑ Mehmedic, Z.; Chandra, S.; Slade, D. et al. (2010). "Potency trends of Δ9-THC and other cannabinoids in confiscated cannabis preparations from 1993 to 2008". Journal of Forensic Sciences 55 (5): 1209–17. doi:10.1111/j.1556-4029.2010.01441.x. PMID 20487147.
- ↑ ElSohly, M.A.; Mehmedic, Z.; Foster, S. et al. (2016). "Changes in Cannabis Potency Over the Last 2 Decades (1995-2014): Analysis of Current Data in the United States". Biological Psychiatry 79 (7): 613-9. doi:10.1016/j.biopsych.2016.01.004. PMC PMC4987131. PMID 26903403. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=PMC4987131.
- ↑ 21.0 21.1 Andre, C.M.; Hausman, J.-F.; Guerriero, G. (2016). "Cannabis sativa: The Plant of the Thousand and One Molecules". Frontiers in Plant Science 7: 19. doi:10.3389/fpls.2016.00019. PMC PMC4740396. PMID 26870049. http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=PMC4740396.
- ↑ 22.0 22.1 Roy, J.-F.; Deckers, C.; Honnold, R. et al. (21 May 2019). "A Sensitive and Robust Workflow to Measure Residual Pesticides and Mycotoxins from the Canadian Target List in Dry Cannabis Flower" (PDF). Agilent Technologies, Inc. https://gcms.cz/labrulez-bucket-strapi-h3hsga3/paper/application-fod-cannabis_5994-0429en-agilent.pdf.
- ↑ Mastovska, K.; Wylie, P.L. (2012). "Evaluation of a new column backflushing set-up in the gas chromatographic-tandem mass spectrometric analysis of pesticide residues in dietary supplements". Journal of Chromatography A 1265: 155–64. doi:10.1016/j.chroma.2012.09.094. PMID 23084487.
- ↑ Wang, M.; Raman, V.; Zhao, J. et al. (2018). "Application of GC/Q-ToF Combined with Advanced Data Mining and Chemometric Tools in the Characterization and Quality Control of Bay Leaves". Planta medica 84 (14): 1045-1054. doi:10.1055/a-0585-5987. PMID 29539646.
- ↑ ElSohly, M.A.; Radwan, M.M.; Gul, W. et al. (2017). "Phytochemistry of Cannabis sativa L". Progress in the Chemistry of Organic Natural Products 103: 1–36. doi:10.1007/978-3-319-45541-9_1. PMID 28120229.
- ↑ 26.0 26.1 Blumberg, L.M.; Klee, M.S. (1998). "Method Translation and Retention Time Locking in Partition GC". Analytical Chemistry 70 (18): 3828–3839. doi:10.1021/ac971141v.
- ↑ Mercer, G.E.; Jones, J.; Olson, M. et al. (July 2015). "A QuEChERS Procedure Focused on Pesticides and Chemical Contaminants that Require Determination by Gas Chromatography". ResearchGate. U.S. Food and Drug Administration. https://www.researchgate.net/publication/317350539.
- ↑ Norli, H.R.; Christiansen, A.; Holen, B. (2010). "Independent evaluation of a commercial deconvolution reporting software for gas chromatography mass spectrometry analysis of pesticide residues in fruits and vegetables". Journal of Chromatography A 1217 (13): 2056-2064. doi:10.1016/j.chroma.2010.01.022.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. Some grammar and punctuation was cleaned up to improve readability. In some cases important information was missing from the references, and that information was added. Nothing else was changed in accordance with the NoDerivatives portion of the license.