Journal:Beyond cannabinoids: Application of NMR-based metabolomics for the assessment of Cannabis sativa L. crop health
| Full article title | Beyond cannabinoids: Application of NMR-based metabolomics for the assessment of Cannabis sativa L. crop health |
|---|---|
| Journal | Frontiers in Plant Science |
| Author(s) | Fernández, Santiago; Castro, Rossina; López-Radcenco, Andrés; Rodriguez, Paula; Carrera, Inés; García-Carnelli, Carlos; Moyna, Guillermo |
| Author affiliation(s) | Universidad de la República - Montevideo, Universidad de la República - Paysandú |
| Primary contact | Email: carlosga at fq dot edu dot uy |
| Editors | Firoentino, Nunzio |
| Year published | 2023 |
| Volume and issue | 14 |
| Article # | 1025932 |
| DOI | 10.3389/fpls.2023.1025932 |
| ISSN | 1664-462X |
| Distribution license | Creative Commons Attribution 4.0 International |
| Website | https://www.frontiersin.org/articles/10.3389/fpls.2023.1025932/full |
| Download | https://www.frontiersin.org/articles/10.3389/fpls.2023.1025932/pdf (PDF) |
Abstract
While Cannabis sativa L. varieties have been traditionally characterized by their major cannabinoid profile, it is now well established that other plant metabolites can also have physiological effects, including minor cannabinoids, terpenes, and flavonoids. Given the multiple applications of Cannabis in the medical field, it is therefore critical to characterize it according to its chemical composition (i.e., its metabolome) and not only its botanical traits. With this in mind, the cannabinoid and metabolomic profiles from inflorescences of two C. sativa varieties with either high Δ9-tetrahydrocannabinolic acid (THCA) or high cannabidiolic acid (CBDA) contents harvested at different times were studied. According to results from high-performance liquid chromatography (HPLC) and nuclear magnetic resonance-based (NMR-based) untargeted metabolomic analyses of organic and aqueous plant material extracts, we show that in addition to expected variations according to cannabinoid profiles, it is possible to distinguish between harvests of the same variety. In particular, it was possible to correlate variations in the metabolome with presence of powdery mildew, leading to the identification of molecular markers associated with this fungal infection in C. sativa.
Keywords: cannabinoids, Cannabis sativa, NMR-based metabolomics, powdery mildew, chemovar
Introduction
Cannabis (Cannabis sativa L.) is a widely distributed plant that has been a source of fiber, food, oil, and medicines for millennia.[1] The first reports regarding its medicinal use date back to the third millennium BC in ancient China.[2] Since then, several cultures have employed Cannabis preparations to treat a variety of ailments.[3] Since our understanding of its therapeutic potential is continuously improving, the interest in Cannabis as a source of compounds for use in medicinal preparations has increased notably.[4] Furthermore, the non-pharmacological uses of the plant and its derivatives, particularly as a source of fibers and as a recreational drug, are well known.
C. sativa produces an important number of secondary metabolites, and more than 500 compounds have been identified in Cannabis so far.[5] In particular, cannabinoids, terpenoids, and flavonoids stand out.[6] Of the three compound classes, cannabinoids are of great relevance given their biological activity and the fact that they are almost exclusively found in this plant.[7][8] Most of the known pharmacological properties are associated to these naturally-occurring compounds, particularly Δ9-tetrahydrocannabinol (THC) and cannabidiol (CBD). These compounds are present mostly as carboxylic acids in the plant—namely Δ9-tetrahydrocannabinolic acid (THCA) and cannabidiolic acid (CBDA)—and are transformed to the neutral molecules through non-enzymatic decarboxylation.[9]
However, the biological effects of C. sativa do not depend only on the levels of THC and CBD.[4] Indeed, nearly 150 phytocannabinoids which can be classified in different structural subclasses have been isolated from this plant[7], and several of these compounds, sometimes referred to as minor cannabinoids, have pharmacological potential.[10] Furthermore, a number of studies suggest that the interaction or synergy between these cannabinoids and other secondary metabolites leads to more pronounced effects than those produced by the isolated compounds.[11][12][13][14][15] In addition, the botanical diversity of C. sativa has resulted in longstanding discussions regarding its taxonomical characterization. It is now generally accepted that the plant is a single species that can be classified in different subspecies and varieties.[16][17] Independent of the names and botanical traits used to refer to it, the chemical composition is what will determine the pharmacological usefulness of the plant. Therefore, several authors stress that the categorization of C. sativa varieties should be done according to its chemical profile[13] and refer to them as chemical varieties or chemovars.[18]
In addition to secondary metabolite content, the quality of a medicinal plant is also given by the absence of contaminants and adulterants. Natural contaminants are usually introduced during cultivation and storage, and they consist of degradation products, microbial contamination, and heavy metals.[19] In particular, fungi can colonize different organs of the plant, causing a reduction in the quality of products created from the infected plant.[20] Furthermore, the consumption of a contaminated product can cause harmful effects on the health of consumers.[21] Microbial infections can also lead to biotic stress, which can alter the secondary metabolite composition.[22]
While the determination of the main cannabinoid content is a required assay for cannabis products that rely on the biological activity of the plant extract, comprehensive knowledge of the profile of other secondary metabolites in these materials is also relevant.[23] Thus, establishing the metabolic fingerprints can provide important information to assure the composition and quality of a given C. sativa chemovar.[24] Variations in this profile can indicate that the material is not appropriate for its intended uses. In particular, the discrimination between chemovars can be conveniently carried out through nuclear magnetic resonance-based (NMR-based) metabolomic analysis.[25] Indeed, a recent review concludes that metabolomics represents an ideal bioanalytical tool that could greatly assist and accelerate cannabis research and development, even coining the term “cannabinomics.”[26]
In the present report we carry out an exhaustive characterization of high-THCA and high-CBDA C. sativa chemovars, both with medicinal potential, through the determination of metabolite profiles complemented with basic information derived from cannabinoid composition. Briefly, our results not only allowed us to clearly distinguish between the two plant varieties, but also to identify a crop with a microbial infection that, interestingly, does not alter the main cannabinoid profile and yield in a relevant way. Furthermore, through metabolomic analysis it was possible to identify variations in specific metabolites that could shed light on infection mechanisms.
Materials and methods
Chemicals and reagents
Analytical grade chloroform and methanol were from Dorwil (Buenos Aires, Argentina), high-performance liquid chromatography-grade (HPLC-grade) acetonitrile was obtained from Carlo Erba (Val-de-Reuil, France), and formic acid, choline (> 98%), deuterochloroform (CDCl3, 99.8%), and deuterium oxide (D2O, 99.9%) were purchased from Merck (Kenilworth, NJ, USA). Betaine (> 98%) was from Thermo Fisher Scientific (Geel, Belgium). Diazepam (> 99.9%) and standard solutions of CBDA, CBD, cannabichromene (CBC), cannabinol (CBN), cannabigerol (CBG), THCA, THC, and Δ8-THC at 1 mg/mL were purchased from Lipomed (Cambridge, MA, USA). Standard solutions of Δ9-tetrahydrocannabivarinic acid (THCVA), Δ9-tetrahydrocannabivarin (THCV), cannabigerolic acid (CBGA), cannabidivarinic acid (CBDVA), cannabidivarin (CBDV) and cannabichromenic acid (CBCA) at 1 mg/mL were purchased from Cerilliant (Round Rock, TX, USA).
Plant material and sample preparation
Cannabis sativa L. female inflorescences were donated by Khiron Life Sciences Uruguay S.A. and employed in compliance with guidelines from a scientific research license granted by the Instituto de Regulación y Control del Cannabis (IRCCA). Plant material from a drug chemotype high in THCA (chemovar A) and a fiber chemotype high in CBDA (chemovar B) were used. Plants were grown under identical indoor conditions, which consisted of temperature and relative humidity ranges of 24-28 °C and 60-70% for the vegetative stage, and 20-26 °C and 55-70% for the flowering stage. Samples were obtained from three different harvests for each chemovar. In the case of chemovar A, these were carried out in March, June, and September 2020, while those for chemovar B harvests were conducted in March, June, and December 2020.
Samples for cannabinoid composition determinations were dried at 35 °C for 24 hours and ground with a manual herb grinder. Once a particle size range from 0.5 to 2 mm was obtained, 500 mg were extracted with a 9:1 methanol-chloroform mixture by dynamic maceration for 30 minutes at room temperature, followed by sonication for 15 minutes. The extract was then filtered and diluted with methanol to a final volume of 50 mL. A 100 μL aliquot of this solution was evaporated under a nitrogen stream and re-dissolved in 100 μL to 1 mL of acetonitrile containing diazepam (10 μg/mL) as an internal standard. The resulting solutions were employed in chromatographic analyses using the conditions detailed below. These extractions were performed in duplicate.
For NMR-based metabolomic analyses, 20 samples per chemotype from each harvest were collected, frozen in liquid nitrogen, ground manually using a glass rod, and stored at -20 °C. Extractions were carried out following a method originally proposed by Choi et al.[25] Briefly, 500 mg of each sample were suspended in 10 mL of a 1:1:2 methanol-water-chloroform mixture, subjected to vortex agitation for 30 seconds, sonicated for one minute, and then centrifuged at 3000 rpm for 20 minutes. Phases were separated and the extraction of the plant material was repeated. Following phase separation, organic solvents were removed in a rotary evaporator under vacuum at 35 °C, while aqueous phases were dried by lyophilization. Two dry extracts per sample were obtained.
HPLC analysis
The main cannabinoid composition was determined using a recently reported method[27], using a Shimadzu DGU-205R Ultra HPLC-PDA system equipped with a Hypersil C18 (150 × 4.6 mm, 3 μm particle size) column and a Hypersil BDS C18 (10 × 4 mm, 3 μm particle size) pre-column (ThermoFisher Scientific, Waltham, MA, USA). Linear calibration curves (r2 > 0.99) for all cannabinoids analyzed were constructed in the range of 1 to 100 μg/mL. The column oven was set to 25 °C, the injection volume was 10 μL, and the total run time was 18 minutes using a solvent gradient composed of 1.0 mM formic acid buffer at pH 3.53 (solvent A) and acetonitrile (solvent B) as mobile phase at 1 mL/min flow. The gradient started with 70% B for seven minutes, then increased to 83.5% B and held constant for one minute, then increased to 88.5% B over a three-minute period, one additional minute to reach 99% B, then kept constant for two minutes followed by two minute to lower B to 70%, and finally held constant for two minutes. This method allowed the separation of the 14 cannabinoids available from a mix of standards.
NMR experiments
Organic extracts were dissolved in 0.7 mL of CDCl3 and aqueous extracts in 0.7 mL of D2O, and then transferred to 5 mm NMR tubes (New Era Enterprises Inc., Vineland, NJ, USA). All NMR spectra were recorded at 25 °C on a Bruker AVANCE III 500 NMR spectrometer operating at 1H and 13C frequencies of 500.13 and 125.76 MHz, respectively, and equipped with a z-gradient TXI probe. A spectral width of 10 KHz, a data size of 32 K, and a total of 64 scans were employed to record 1D spectra, using a relaxation delay of four seconds between scans. Regular 1D acquisition sequences with 30° excitation pulses were employed, using a standard water pre-saturation scheme with aqueous samples. When required, 1D-TOCSY, 1D-NOESY, HSQC, and HMBC spectra were acquired and processed using acquisition and processing parameters provided with the spectrometer.
To confirm the presence of choline and betaine in the aqueous extracts, 10 μL aliquots from 50 mM standard solutions of each compound were added to selected samples, and the 1D 1H NMR experiments repeated under the same conditions described above.
NMR data processing
NMR data for metabolomic analyses were processed and analyzed with MNova (version 12.0, MestreLab Research, S.L., Santiago de Compostela, Spain). Free induction decays were zero-filled to 64 K points and apodized with a 0.3 Hz exponential window function prior to Fourier transformation. All spectra were manually phase- and baseline-corrected and referenced. In the case of organic extracts, the residual CHCl3 solvent signal at 7.28 ppm was used for calibration, while the anomeric proton signal of α-glucose at 5.22 ppm was employed in aqueous samples. Spectra corresponding to the same class of extract were aligned using PAFFT[28], and the data was normalized to the total spectral area after excluding residual solvent resonances and regions without signals. The resulting data matrices were finally exported as text files for use in statistical analyses.
Multivariate statistical analysis
Multivariate statistical analyses, including principal component analysis (PCA) and orthogonal partial least squares discriminant analysis (OPLS-DA), were carried out with the PLS_Toolbox package (version 8.5, Eigenvector Research Inc., Manson, WA, USA) implemented for MATLAB (revision 2014a, The MathWorks Inc., Natick, MA, USA). For all models, the data was mean-centered and scaled using a Pareto factor.[29] Analysis of the data was first performed with PCA, which reduces the dimensionality and facilitates the identification of data clusters or trends.[30][31][32] The PCA scores plot was also employed to identify strong outliers outside the 95% significance region of Hotelling’s T2 ellipse. Cross-validation of OPLS-DA models was achieved using the random subset method, which involved 20 iterations over data split into 10 equally-sized parts. Receiver operating characteristic (ROC) curves were plotted, and areas under the curves were calculated to ensure the goodness of fit of the resulting models.[33][34] Permutation tests with 100 iterations were also performed to determine the degree of over-fitting and further validate the discriminant analyses.[35] When needed, statistical total correlation spectroscopy (STOCSY) analyses were performed with an in-house MATLAB script based on the algorithm described by Cloarec et al.[36]
Fungi characterization
The fungal infection was evidenced by the presence of off-white powdery spots typical of powdery mildew disease. Microscopic examination of fungi was carried out with a Nikon Eclipse E100 optical microscope. To examine asexual morphs, a piece of clear adhesive tape was placed on infected leaves, stripped off, and placed on a microscope slide with one drop of distilled water. The observations were done at magnifications of 40, 100, and 400 under standard light. Germinated conidia were examined on fungi grown with potato dextrose agar medium.
Results
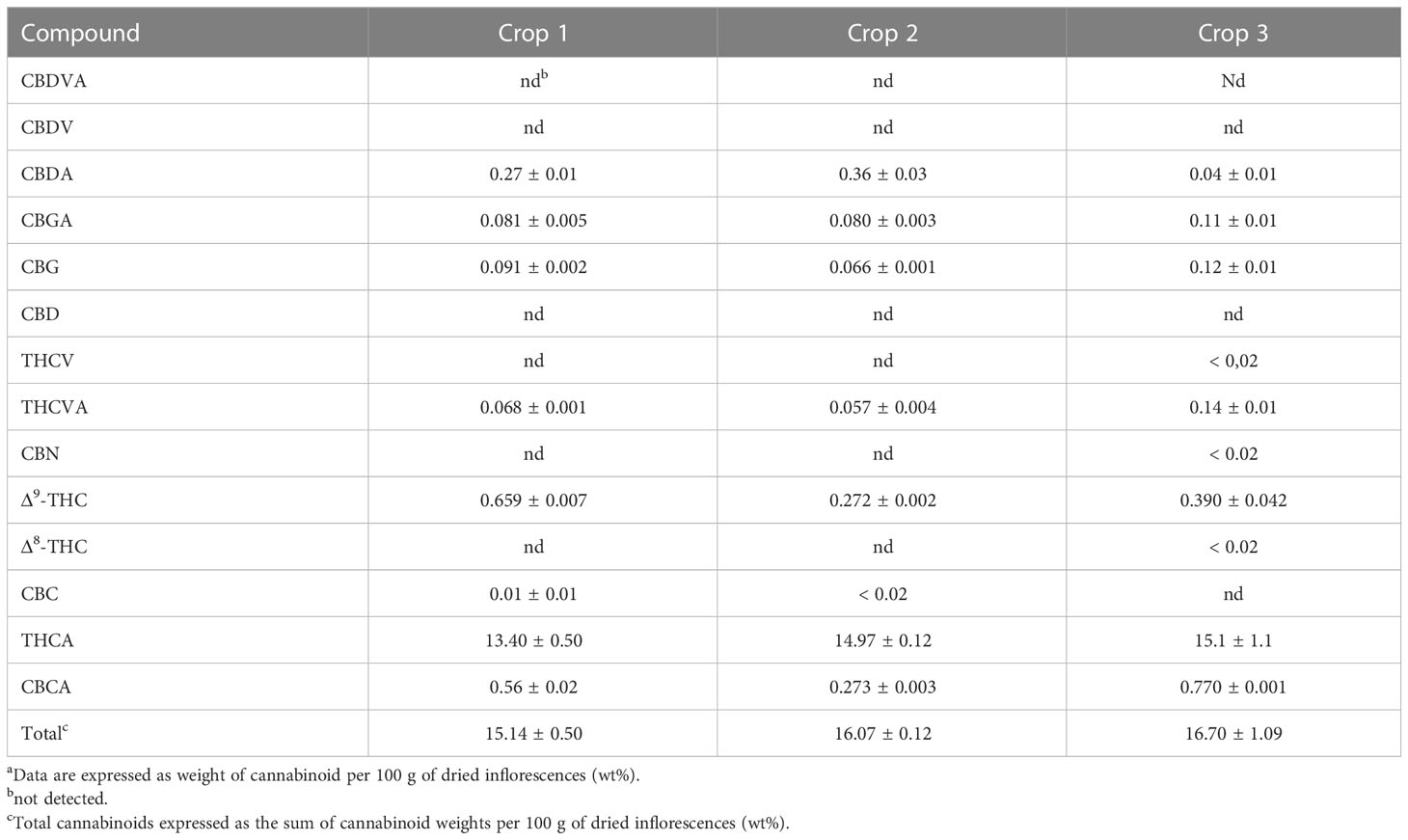
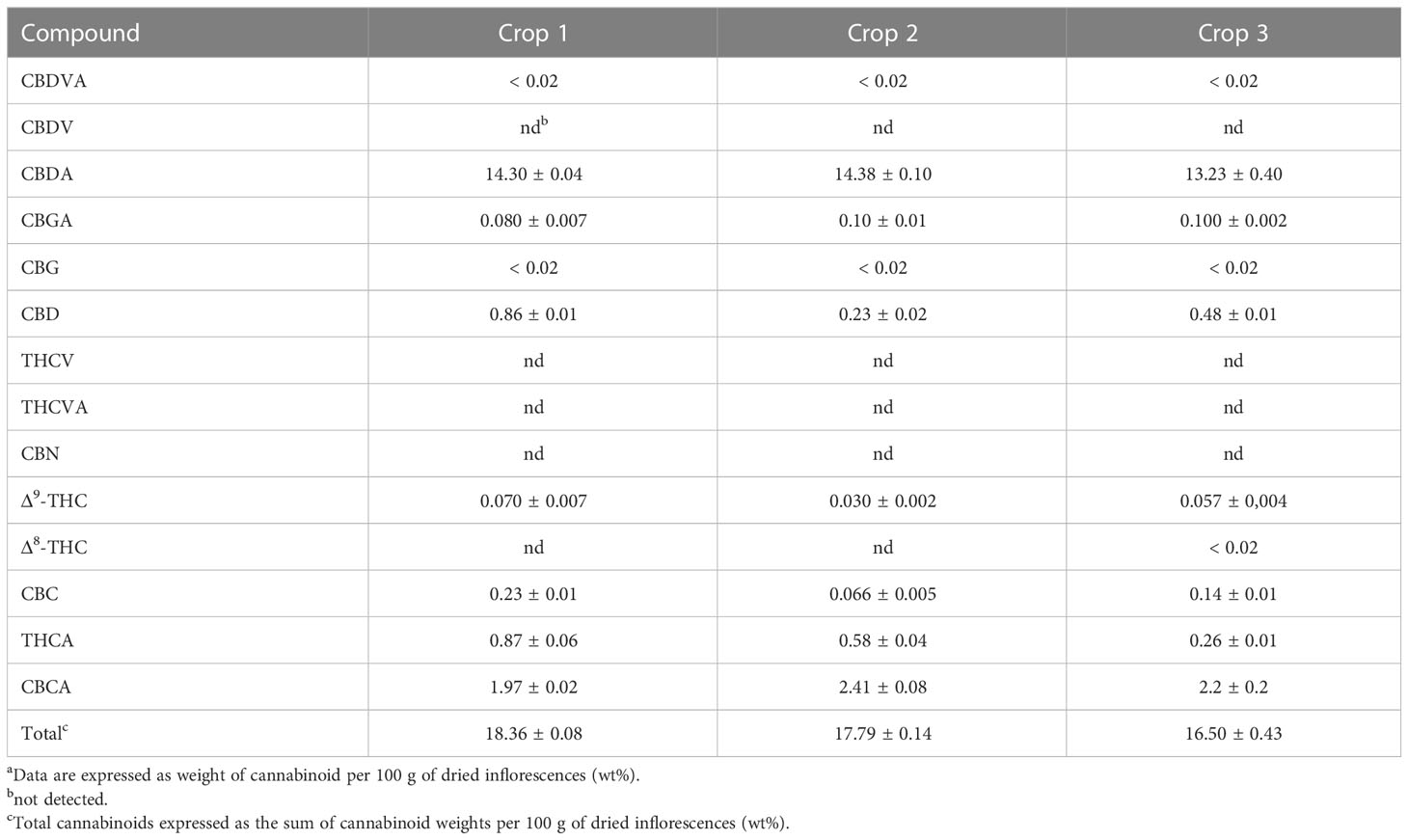
Tables 1 and 2 show the cannabinoid composition of chemovars A and B, respectively. As expected, the two are notably different, and characterized, respectively, by their particularly high content of THCA and CBDA. Moreover, the acidic forms of other cannabinoids are also predominant, and CBN was only detected in one sample and below the quantification limits. This suggests that little degradation occurred during sample handling and analysis. For chemovar A, the total cannabinoid content ranged from 15.14 ± 0.50 to 16.70 ± 1.10%, with a mean of 15.50 ± 1.10% (n = 6, two samples per crop, three crops). Similarly, the cannabinoid content in chemovar B crops ranged from 16.50 ± 0.43 to 18.36 ± 0.08% with a mean of 17.60 ± 0.80% (n = 6, two samples per crop, three crops). These results agree with literature reports, which indicate average cannabinoid contents in inflorescences ranging from 5 to 25%.[23] Inspection of the results also shows that relative cannabinoid composition and total cannabinoid yield is stable through the crops in both chemovars.
|
|
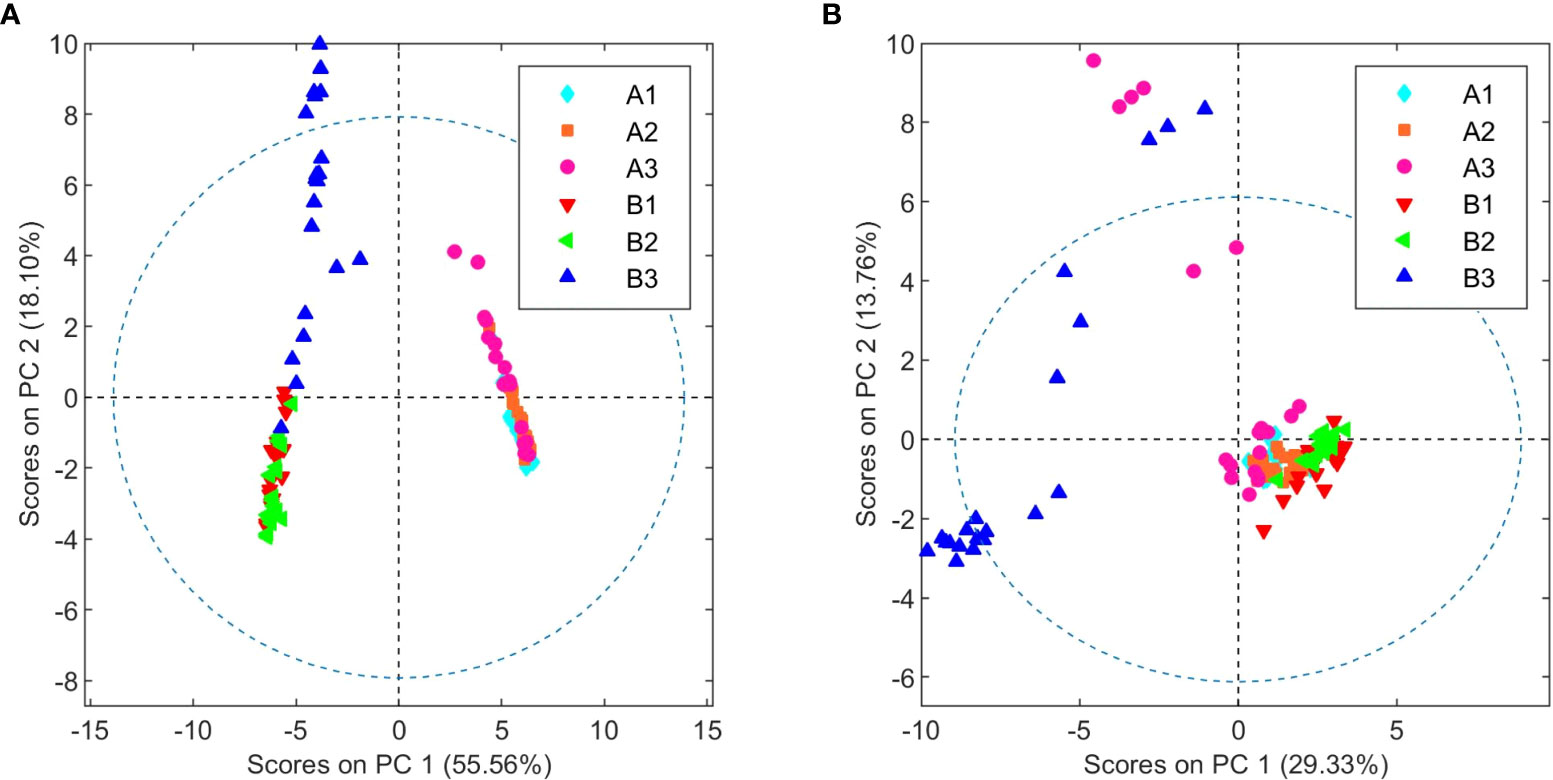
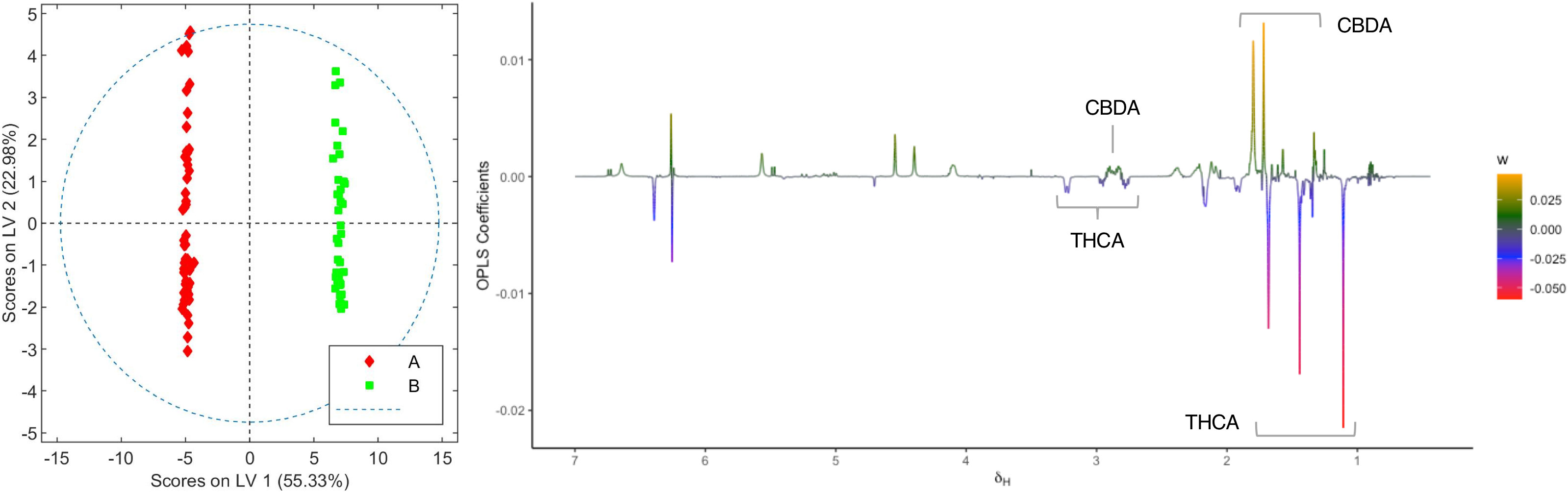
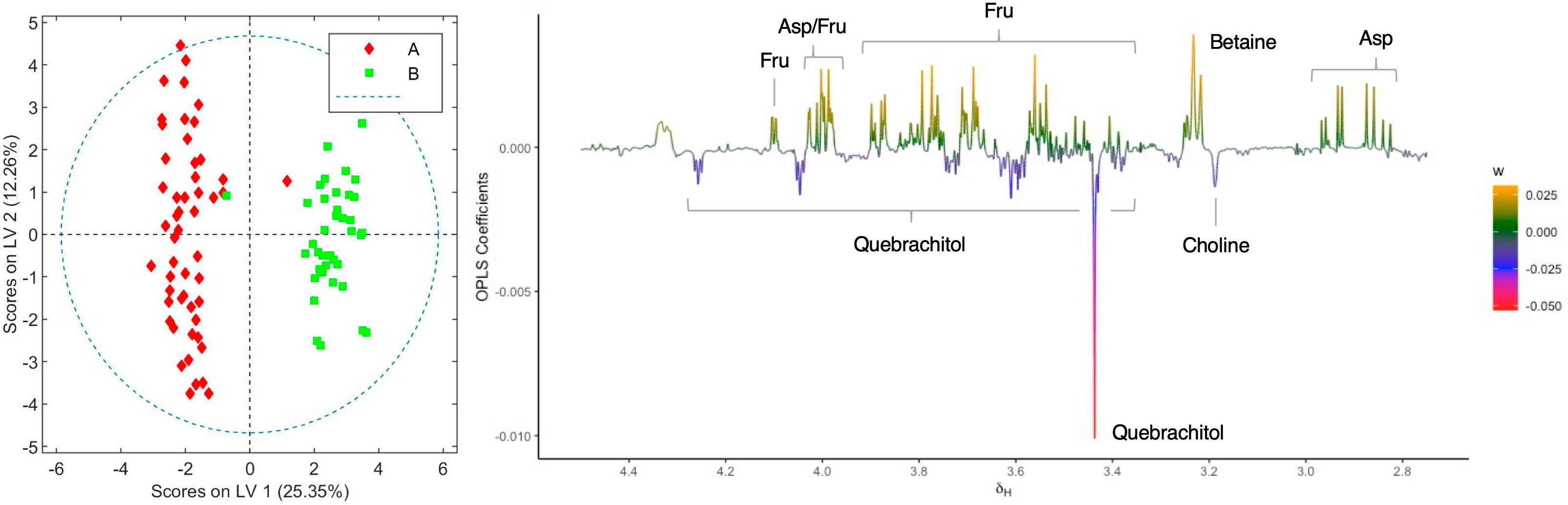
Following the determination of the basic cannabinoid composition, the metabolic profiles for both C. sativa varieties were investigated. For this purpose, 20 samples of each chemovar from the three harvests considered were collected, resulting in a total of 60 organic and 60 aqueous extract samples. PCA score plots obtained from the 1H NMR data recorded for the two sets are shown in Figure 1. It is clear that both chemovars can be easily differentiated on the basis of NMR data from their organic or aqueous extracts. Indeed, the first two principal components (PCs) account for 73.7% of the variance in the organic set, with samples from chemovar A forming a cluster showing positive PC1 scores, whereas chemovar B samples have negative scores on this PC. Similarly, only two PCs were sufficient to account for nearly 90% of the variance in the aqueous data set.
|
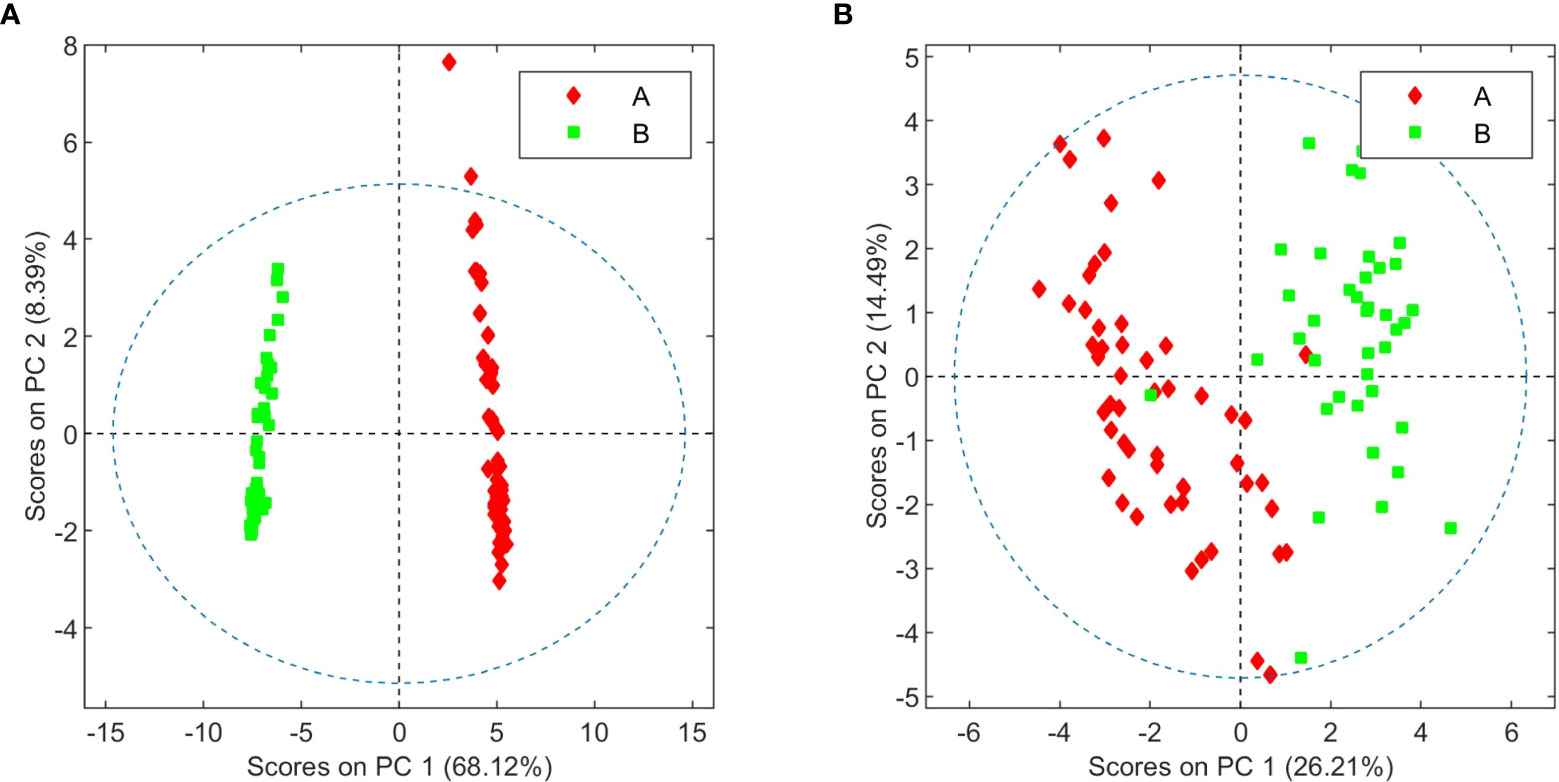
Cursory inspection of both PCA plots indicates that six of the aqueous samples corresponding to the third crop of chemovar A deviate from the others, suggesting errors during the preparation of these samples. On the other hand, all 20 organic and aqueous extracts from the third crop of chemovar B deviate considerably from the rest. As discussed below, this is likely associated with fungal infections affecting plants from this particular crop. Therefore, the PCA score plots were recomputed without the outliers from both chemovars. As shown in Figure 2, 1H NMR profiles from organic and aqueous extracts can still differentiate the two chemovars clearly.
|
In order to identify the metabolites that differentiate the two chemovars, OPLS-DA models from the depurated organic and aqueous extract 1H NMR datasets were generated. Inspection of the OPLS-DA loading factor plot obtained from organic extracts indicates that signals at 1.44, 1.25, and 1.11 ppm, corresponding to THCA, correlate to chemovar A, while signals for CBDA at 1.80 and 1.72 ppm corroborate that the levels of this cannabinoid are higher in chemovar B (Figure 3).[37] These results are not surprising and are consistent with those obtained by the HPLC analyses presented in Tables 1 and 2.
|
When the loading factor plot of the OLPS-DA between aqueous extracts of the two chemovars was analyzed (Figure 4), positive signals at 4.10, 4.03–3.97, 3.88, 3.82–3.77, 3.70–3.63, and 3.59–3.52 ppm, and at 3.99, 2.93, and 2.84 ppm, which can be respectively assigned to fructose and aspartate by comparison to literature data[38][39][40], show that the levels of these two metabolites are higher in chemovar B plants. In addition, a positive signal at 3.23 ppm with an HSQC correlation to a 13C resonance at 53.4 ppm likely corresponds to the N-methyl groups of betaine[40][41], indicating that the levels of this trimethylammonium-bearing metabolite are also higher in the high-CBDA chemovar specimens (Supplementary material, Figure S3). This initial assignment was confirmed by spiking one of the aqueous extract samples with a betaine standard solution (see Supplementary material, Figure S4). On the other hand, negative peaks at 4.24, 4.04, 3.73, 3.59, 3.57, 3.44, and 3.38 ppm match nicely with reference data for the carbasugar quebrachitol.[42] This reveals that the concentration of this cyclitol, whose identity was further established by means of STOCSY, 1D-TOCSY, 1D-NOESY, and heteronuclear correlation NMR experiments (see Supplementary material, Figures S5–S7; Table S1), is higher in chemovar A plants. Moreover, a negative singlet at 3.18 ppm correlating to a 13C resonance at 53.9 ppm in the HSQC spectrum corresponds to choline[40][41], suggesting that the levels of this quaternary aminoalcohol are higher in the high-THCA C. sativa variety (Supplementary material, Figure S8). As was the case for betaine, this assignment was confirmed unequivocally by spiking experiments (Supplementary material, Figure S4).
|
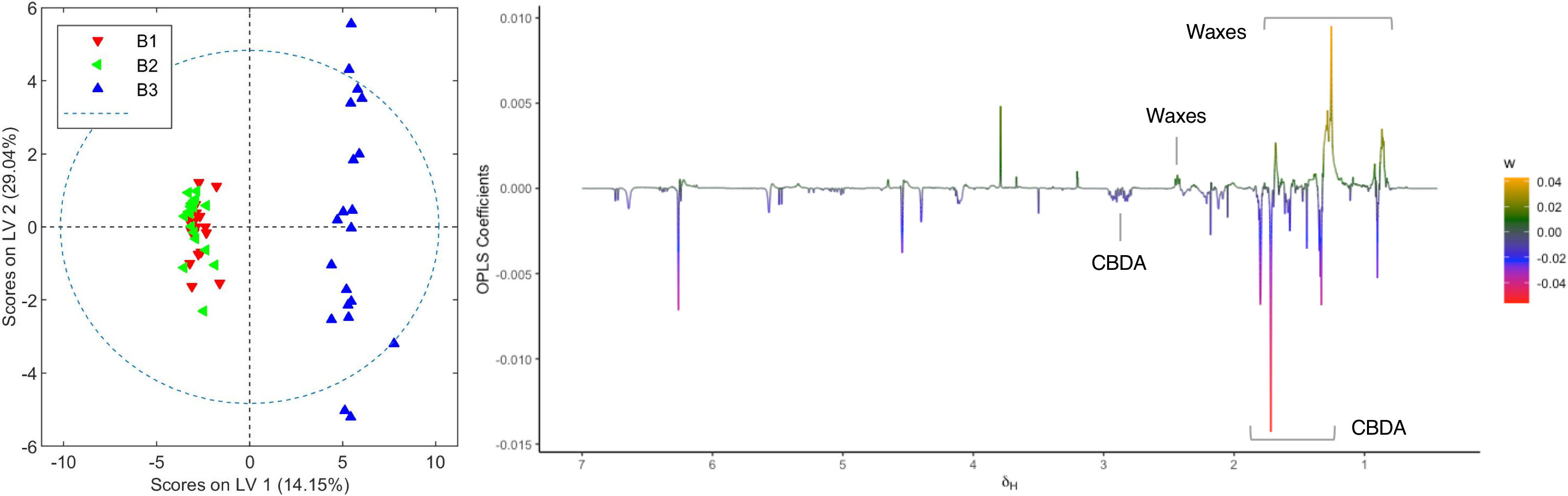
Plants from the third crop of chemovar B showed signs of powdery mildew infection. The morphological characteristics of this fungal leaf growth and its asexual microscopic structures allowed us to classify it as a member of the Golovinomyces genus (Supplementary material, Figures S11 and S12). Since this infection was also clearly evidenced in the initial clustering analyses depicted in Figure 1, we decided to carry out a more exhaustive comparison between healthy and infected plants within this chemovar using OPLS-DA. As shown in Figure 5, the model obtained with 1H NMR data from organic extracts easily differentiates inflorescences of the first two crops (B1+B2) from those of the third (B3). The corresponding loading factor plot identifies CBDA as one of the discriminating metabolites in this case, and, in agreement with HPLC findings presented in Table 2, indicates that higher concentrations of this cannabinoid are present in healthy plants of the high-CBDA chemovar. On the other hand, higher levels of waxes, with signals at 2.42, 1.70, 1.28–1.25, and 0.92 ppm[43], are found in infected plant extracts.
|
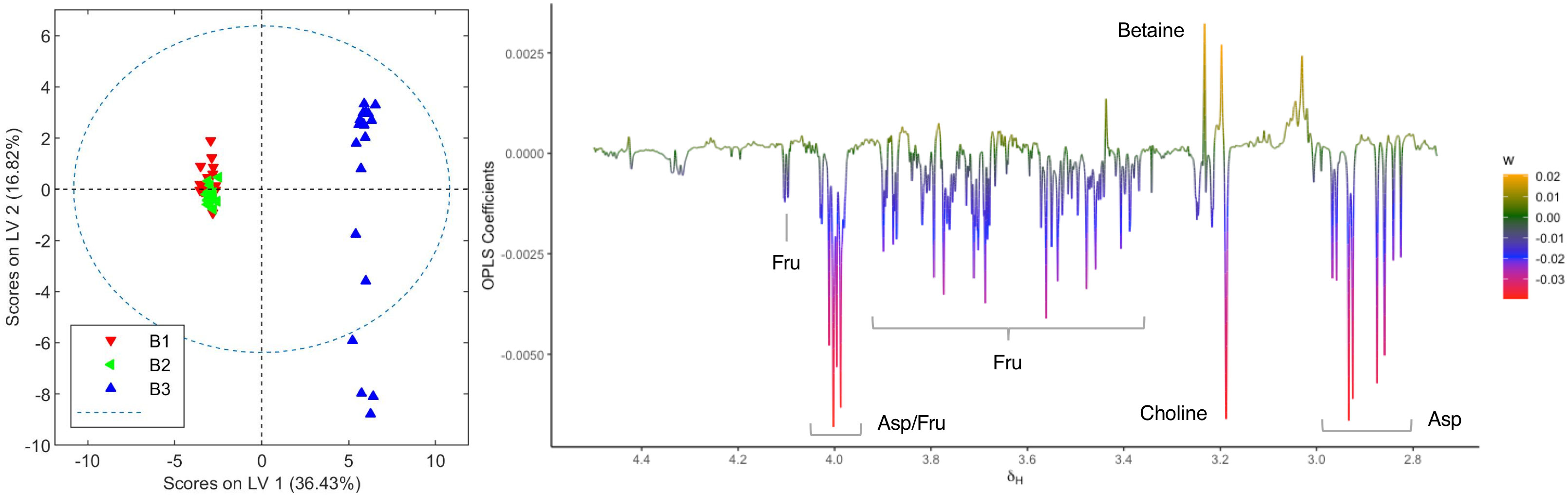
The OPLS-DA model comparing aqueous extracts from the B1+B2 harvest versus those from B3 can also classify healthy and infected plants correctly (Figure 6). As observed in the loading factor plot, aspartate, choline, and fructose are in this case associated with healthy chemovar B specimens, while the levels of betaine are higher in infected plants.
|
Discussion
As stated in the introduction, our initial goal was to perform an exhaustive chemical characterization of high-THCA and high-CBDA C. sativa chemovars with medicinal potential through the determination of their metabolic profiles. Our results revealed a number of differences between the two varieties beyond their cannabinoid composition. This is an important finding since more than 500 different compounds have been reported in the plant, some with potential therapeutic qualities, and combinations of the various secondary metabolites could determine both the final medicinal response as well as any adverse effects.[4][44] In fact—and despite the results from NMR-based metabolomic analysis of organic extracts presented in Figures 2 and 3 paralleling those obtained through HPLC (Tables 1 and 2)—the study of aqueous fractions identified aspartate, betaine, choline, fructose, and quebrachitol as metabolites that can further discriminate between the two chemovars. The differences in sugar and aminoacid contents, which have been described by Choi et al. in their seminal report on the application of metabolomic profiling to the characterization of cannabis cultivars[25], could be employed to better fingerprint cannabis varieties and products intended for use as botanical drugs. This is also the case for quebrachitol, a long-known component of C. sativa.[45][46] Indeed, variations in the concentration of this cyclitol have also been employed to address the phenotypical plasticity in other plants[47], making it a promising phytomarker for cultivar differentiation. Given the correlation between betaine levels and abiotic stress[48], the identification of higher concentrations of this metabolite in high-CBDA chemovars suggests better adaptation to drought, high salinity, and low temperature conditions. In principle, this information would be instrumental to breeders in the development of plant varieties agronomically adapted to specific environmental conditions.
In addition to differences between varieties, analysis of the data from different harvests also allowed us to identify changes in the metabolome caused by fungal infection. As summarized in Figures 5 and 6, higher levels of CBDA, fructose, aspartate, and choline were observed in healthy plants from chemovar B, while increased levels of waxes and betaine were found in infected specimens. It is well known that biotic and abiotic stresses, and in particular bacterial and fungal infections, can lead to variations in the cannabinoid profile in C. sativa.[22] While these variations were not evident when comparing the cannabinoid compositions of the three crops using data from standard HPLC methods, multivariate analysis of NMR data was highly sensitive and allowed to readily detect them. Indeed, CBDA was identified as one of the metabolites that can aid in the classification of healthy and diseased plants. While HPLC revealed a slightly lower concentration for this metabolite in the third crop relative to the first two (13.23% versus 14.30 and 14.38%), the variation is well within quality specifications proposed for botanical drugs.[49] Furthermore, the full cannabinoidic profile obtained by HPLC was the same for the three crops. In agreement with their roles in stress response, the concentration of waxes and betaine was also higher in plants with signs of infection. The former compounds constitute one of the first lines of defense against microbes.[50][51] While larger alterations in wax composition could be expected in leaves, variations in its concentration in inflorescences are also likely upon fungal infection[52] Similarly, and taking into account its role as an osmoprotectant that counteracts the effects of reactive oxygen species and promotes membrane stabilization[48], variations in betaine levels are consistent with the necrotic lesions observed on plant tissue affected by powdery mildew.[20] As discussed above, variations in betaine concentration in plants are normally associated with abiotic stress. However, there is evidence that fungal infections can also lead to increases in the levels of this quaternary ammonium alcohol in other crops.[53][54]
To summarize, our studies contribute to the holistic chemical characterization of C. sativa varieties with potential medicinal applications. Given the variability in the chemical composition of these plants and their extracts, this information is critical to assure the quality of botanical drugs and related products derived from them. In addition, the results presented above further showcase the suitability of metabolomic profiling as a tool for the rational classification of plant materials.
Supplementary material
The supplementary material for this article can be found here: Data Sheet 1.docx
Acknowledgements
The authors wish to thank Drs. Agustina Vila and Victoria Giorgi for insightful suggestions and technical assistance. We are also grateful to Prof. Dr. Patrick Moyna for reviewing the grammar and wording of the final draft.
Author contributions
SF: methodology, data curation, formal analysis, investigation, and writing–original draft. RC: data curation and formal analysis. AL-R: data curation, formal analysis, and investigation. PR: data curation, formal analysis, and investigation. IC: project administration, funding acquisition, and writing–review and editing. CG-C: conceptualization, methodology, project administration, funding acquisition, and writing–review and editing. GM: conceptualization, methodology, project administration, and writing-review and editing. All authors contributed to the article and approved the submitted version.
Funding
This work was jointly funded by Agencia Nacional de Investigación e Innovación (ANII) and Khiron Life Sciences Uruguay S.A. (award ALI_1_2018_1_147904). Additional financial support was received from the Programa para el Desarrollo de las Ciencias Básicas (PEDECIBA).
Data availability statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The authors declare that this study received funding from Khiron Life Sciences Uruguay S.A. The funder was not involved in the study design, collection, analysis, interpretation of data, the writing of this article or the decision to submit it for publication.
References
- ↑ Clarke, Robert C.; Merlin, Mark D. (1 November 2016). "Cannabis Domestication, Breeding History, Present-day Genetic Diversity, and Future Prospects" (in en). Critical Reviews in Plant Sciences 35 (5-6): 293–327. doi:10.1080/07352689.2016.1267498. ISSN 0735-2689. https://www.tandfonline.com/doi/full/10.1080/07352689.2016.1267498.
- ↑ Wills, S. (1998). "Cannabis Use and Abuse by Man: An Historical Perspective". In Brown, D.T.. Cannabis: The Genus Cannabis (1st ed.). CRC Press. pp. 1–28. ISBN 9780429219771.
- ↑ Russo, Ethan B. (21 August 2014), Pertwee, Roger, ed., "The Pharmacological History of Cannabis", Handbook of Cannabis (Oxford University Press): 23–43, doi:10.1093/acprof:oso/9780199662685.003.0002, ISBN 978-0-19-966268-5, https://academic.oup.com/book/27329/chapter/197025364. Retrieved 2023-04-12
- ↑ 4.0 4.1 4.2 Procaccia, Shiri; Lewitus, Gil Moshe; Lipson Feder, Carni; Shapira, Anna; Berman, Paula; Meiri, David (25 April 2022). "Cannabis for Medical Use: Versatile Plant Rather Than a Single Drug". Frontiers in Pharmacology 13: 894960. doi:10.3389/fphar.2022.894960. ISSN 1663-9812. PMC PMC9081504. PMID 35548332. https://www.frontiersin.org/articles/10.3389/fphar.2022.894960/full.
- ↑ ElSohly, Mahmoud A.; Gul, Waseem (21 August 2014), Pertwee, Roger, ed., "Constituents of Cannabis Sativa", Handbook of Cannabis (Oxford University Press): 3–22, doi:10.1093/acprof:oso/9780199662685.003.0001, ISBN 978-0-19-966268-5, https://academic.oup.com/book/27329/chapter/197023611
- ↑ ElSohly, Mahmoud A.; Slade, Desmond (1 December 2005). "Chemical constituents of marijuana: The complex mixture of natural cannabinoids" (in en). Life Sciences 78 (5): 539–548. doi:10.1016/j.lfs.2005.09.011. https://linkinghub.elsevier.com/retrieve/pii/S002432050500891X.
- ↑ 7.0 7.1 Hanuš, Lumír Ondřej; Meyer, Stefan Martin; Muñoz, Eduardo; Taglialatela-Scafati, Orazio; Appendino, Giovanni (2016). "Phytocannabinoids: a unified critical inventory" (in en). Natural Product Reports 33 (12): 1357–1392. doi:10.1039/C6NP00074F. ISSN 0265-0568. http://xlink.rsc.org/?DOI=C6NP00074F.
- ↑ Gülck, Thies; Møller, Birger Lindberg (1 October 2020). "Phytocannabinoids: Origins and Biosynthesis" (in en). Trends in Plant Science 25 (10): 985–1004. doi:10.1016/j.tplants.2020.05.005. https://linkinghub.elsevier.com/retrieve/pii/S1360138520301874.
- ↑ Flores-Sanchez, Isvett Josefina; Verpoorte, Robert (1 December 2008). "PKS Activities and Biosynthesis of Cannabinoids and Flavonoids in Cannabis sativa L. Plants" (in en). Plant and Cell Physiology 49 (12): 1767–1782. doi:10.1093/pcp/pcn150. ISSN 1471-9053. https://academic.oup.com/pcp/article-lookup/doi/10.1093/pcp/pcn150.
- ↑ Franco, Rafael; Rivas-Santisteban, Rafael; Reyes-Resina, Irene; Casanovas, Mireia; Pérez-Olives, Catalina; Ferreiro-Vera, Carlos; Navarro, Gemma; Sánchez de Medina, Verónica et al. (1 August 2020). "Pharmacological potential of varinic-, minor-, and acidic phytocannabinoids" (in en). Pharmacological Research 158: 104801. doi:10.1016/j.phrs.2020.104801. https://linkinghub.elsevier.com/retrieve/pii/S1043661820311099.
- ↑ Russo, Ethan B (1 August 2011). "Taming THC: potential cannabis synergy and phytocannabinoid-terpenoid entourage effects: Phytocannabinoid-terpenoid entourage effects" (in en). British Journal of Pharmacology 163 (7): 1344–1364. doi:10.1111/j.1476-5381.2011.01238.x. PMC PMC3165946. PMID 21749363. https://onlinelibrary.wiley.com/doi/10.1111/j.1476-5381.2011.01238.x.
- ↑ Blasco-Benito, Sandra; Seijo-Vila, Marta; Caro-Villalobos, Miriam; Tundidor, Isabel; Andradas, Clara; García-Taboada, Elena; Wade, Jeff; Smith, Stewart et al. (1 November 2018). "Appraising the “entourage effect”: Antitumor action of a pure cannabinoid versus a botanical drug preparation in preclinical models of breast cancer" (in en). Biochemical Pharmacology 157: 285–293. doi:10.1016/j.bcp.2018.06.025. https://linkinghub.elsevier.com/retrieve/pii/S0006295218302387.
- ↑ 13.0 13.1 Russo, Ethan B. (9 January 2019). "The Case for the Entourage Effect and Conventional Breeding of Clinical Cannabis: No “Strain,” No Gain". Frontiers in Plant Science 9: 1969. doi:10.3389/fpls.2018.01969. ISSN 1664-462X. PMC PMC6334252. PMID 30687364. https://www.frontiersin.org/article/10.3389/fpls.2018.01969/full.
- ↑ Maayah, Zaid H.; Takahara, Shingo; Ferdaoussi, Mourad; Dyck, Jason R. B. (1 June 2020). "The anti-inflammatory and analgesic effects of formulated full-spectrum cannabis extract in the treatment of neuropathic pain associated with multiple sclerosis" (in en). Inflammation Research 69 (6): 549–558. doi:10.1007/s00011-020-01341-1. ISSN 1023-3830. http://link.springer.com/10.1007/s00011-020-01341-1.
- ↑ Goerl, Brett; Watkins, Sarah; Metcalf, Cameron; Smith, Misty; Beenhakker, Mark (1 January 2021). "Cannabidiolic acid exhibits entourage-like improvements of anticonvulsant activity in an acute rat model of seizures" (in en). Epilepsy Research 169: 106525. doi:10.1016/j.eplepsyres.2020.106525. PMC PMC7855831. PMID 33310415. https://linkinghub.elsevier.com/retrieve/pii/S0920121120305763.
- ↑ Small, Ernest (1 September 2015). "Evolution and Classification of Cannabis sativa (Marijuana, Hemp) in Relation to Human Utilization" (in en). The Botanical Review 81 (3): 189–294. doi:10.1007/s12229-015-9157-3. ISSN 0006-8101. http://link.springer.com/10.1007/s12229-015-9157-3.
- ↑ McPartland, John M. (1 October 2018). "Cannabis Systematics at the Levels of Family, Genus, and Species" (in en). Cannabis and Cannabinoid Research 3 (1): 203–212. doi:10.1089/can.2018.0039. ISSN 2378-8763. PMC PMC6225593. PMID 30426073. https://www.liebertpub.com/doi/10.1089/can.2018.0039.
- ↑ Lewis, Mark; Russo, Ethan; Smith, Kevin (1 March 2018). "Pharmacological Foundations of Cannabis Chemovars" (in en). Planta Medica 84 (04): 225–233. doi:10.1055/s-0043-122240. ISSN 0032-0943. http://www.thieme-connect.de/DOI/DOI?10.1055/s-0043-122240.
- ↑ Cannabis Inflorescence: Standards of identity, analysis and quality control (Revision 2014 ed.). American Herbal Pharmacopoeia. 2014. ISBN 1929425376. ISSN 1538-0297. https://herbal-ahp.com/collections/frontpage/products/cannabis-inflorescence-quality-control-monograph.
- ↑ 20.0 20.1 Punja, Zamir K.; Collyer, Danielle; Scott, Cameron; Lung, Samantha; Holmes, Janesse; Sutton, Darren (17 October 2019). "Pathogens and Molds Affecting Production and Quality of Cannabis sativa L.". Frontiers in Plant Science 10: 1120. doi:10.3389/fpls.2019.01120. ISSN 1664-462X. PMC PMC6811654. PMID 31681341. https://www.frontiersin.org/article/10.3389/fpls.2019.01120/full.
- ↑ Montoya, Zackary; Conroy, Matthieu; Vanden Heuvel, Brian D.; Pauli, Christopher S.; Park, Sang-Hyuck (11 September 2020). "Cannabis Contaminants Limit Pharmacological Use of Cannabidiol". Frontiers in Pharmacology 11: 571832. doi:10.3389/fphar.2020.571832. ISSN 1663-9812. PMC PMC7516211. PMID 33013414. https://www.frontiersin.org/article/10.3389/fphar.2020.571832/full.
- ↑ 22.0 22.1 Gorelick, Jonathan; Bernstein, Nirit (2017), Chandra, Suman; Lata, Hemant; ElSohly, Mahmoud A., eds., "Chemical and Physical Elicitation for Enhanced Cannabinoid Production in Cannabis" (in en), Cannabis sativa L. - Botany and Biotechnology (Cham: Springer International Publishing): 439–456, doi:10.1007/978-3-319-54564-6_21, ISBN 978-3-319-54563-9, http://link.springer.com/10.1007/978-3-319-54564-6_21. Retrieved 2023-04-12
- ↑ 23.0 23.1 Jin, Dan; Henry, Philippe; Shan, Jacqueline; Chen, Jie (1 July 2021). "Identification of Chemotypic Markers in Three Chemotype Categories of Cannabis Using Secondary Metabolites Profiled in Inflorescences, Leaves, Stem Bark, and Roots". Frontiers in Plant Science 12: 699530. doi:10.3389/fpls.2021.699530. ISSN 1664-462X. PMC PMC8283674. PMID 34276749. https://www.frontiersin.org/articles/10.3389/fpls.2021.699530/full.
- ↑ Puri, Shivani; Sahal, Dinkar; Sharma, Upendra (1 December 2021). "A conversation between hyphenated spectroscopic techniques and phytometabolites from medicinal plants" (in en). Analytical Science Advances 2 (11-12): 579–593. doi:10.1002/ansa.202100021. ISSN 2628-5452. https://chemistry-europe.onlinelibrary.wiley.com/doi/10.1002/ansa.202100021.
- ↑ 25.0 25.1 25.2 Choi, Young Hae; Hazekamp, Arno; Peltenburg-Looman, Anja M. G.; Frédérich, Michel; Erkelens, Cornelis; Lefeber, Alfons W. M.; Verpoorte, Robert (1 November 2004). "NMR assignments of the major cannabinoids and cannabiflavonoids isolated from flowers of Cannabis sativa: NMR ASSIGNMENTS OF CANNABINOIDS AND CANNABIFLAVONOIDS" (in en). Phytochemical Analysis 15 (6): 345–354. doi:10.1002/pca.787. https://onlinelibrary.wiley.com/doi/10.1002/pca.787.
- ↑ Aliferis, Konstantinos A.; Bernard-Perron, David (8 May 2020). "Cannabinomics: Application of Metabolomics in Cannabis (Cannabis sativa L.) Research and Development". Frontiers in Plant Science 11: 554. doi:10.3389/fpls.2020.00554. ISSN 1664-462X. PMC PMC7225349. PMID 32457786. https://www.frontiersin.org/article/10.3389/fpls.2020.00554/full.
- ↑ Fernández, Santiago; Carreras, Tatiana; Castro, Rossina; Perelmuter, Karen; Giorgi, Victoria; Vila, Agustina; Rosales, Agustina; Pazos, Mariana et al. (1 January 2022). "A comparative study of supercritical fluid and ethanol extracts of cannabis inflorescences: Chemical profile and biological activity" (in en). The Journal of Supercritical Fluids 179: 105385. doi:10.1016/j.supflu.2021.105385. https://linkinghub.elsevier.com/retrieve/pii/S0896844621002278.
- ↑ Wong, Jason W. H.; Durante, Caterina; Cartwright, Hugh M. (1 September 2005). "Application of Fast Fourier Transform Cross-Correlation for the Alignment of Large Chromatographic and Spectral Datasets" (in en). Analytical Chemistry 77 (17): 5655–5661. doi:10.1021/ac050619p. ISSN 0003-2700. https://pubs.acs.org/doi/10.1021/ac050619p.
- ↑ van den Berg, Robert A; Hoefsloot, Huub CJ; Westerhuis, Johan A; Smilde, Age K; van der Werf, Mariët J (1 December 2006). "Centering, scaling, and transformations: improving the biological information content of metabolomics data" (in en). BMC Genomics 7 (1): 142. doi:10.1186/1471-2164-7-142. ISSN 1471-2164. PMC PMC1534033. PMID 16762068. https://bmcgenomics.biomedcentral.com/articles/10.1186/1471-2164-7-142.
- ↑ Wold, Svante; Esbensen, Kim; Geladi, Paul (1 August 1987). "Principal component analysis" (in en). Chemometrics and Intelligent Laboratory Systems 2 (1-3): 37–52. doi:10.1016/0169-7439(87)80084-9. https://linkinghub.elsevier.com/retrieve/pii/0169743987800849.
- ↑ Trygg, Johan; Wold, Svante (1 March 2002). "Orthogonal projections to latent structures (O-PLS)" (in en). Journal of Chemometrics 16 (3): 119–128. doi:10.1002/cem.695. ISSN 0886-9383. https://onlinelibrary.wiley.com/doi/10.1002/cem.695.
- ↑ Trygg, Johan; Holmes, Elaine; Lundstedt, Torbjörn (1 February 2007). "Chemometrics in Metabonomics" (in en). Journal of Proteome Research 6 (2): 469–479. doi:10.1021/pr060594q. ISSN 1535-3893. https://pubs.acs.org/doi/10.1021/pr060594q.
- ↑ Ekelund, Suzanne (1 March 2012). "ROC Curves—What are They and How are They Used?" (in en). Point of Care: The Journal of Near-Patient Testing & Technology 11 (1): 16–21. doi:10.1097/POC.0b013e318246a642. ISSN 1533-029X. https://journals.lww.com/00134384-201203000-00006.
- ↑ Simundic, Ana-Maria (2012). "Practical recommendations for statistical analysis and data presentation in Biochemia Medica journal". Biochemia Medica: 15–23. doi:10.11613/BM.2012.003. http://www.biochemia-medica.com/en/journal/22/1/10.11613/BM.2012.003.
- ↑ Ni, Yan; Su, Mingming; Lin, Jinchao; Wang, Xiaoyan; Qiu, Yunping; Zhao, Aihua; Chen, Tianlu; Jia, Wei (23 July 2008). "Metabolic profiling reveals disorder of amino acid metabolism in four brain regions from a rat model of chronic unpredictable mild stress" (in en). FEBS Letters 582 (17): 2627–2636. doi:10.1016/j.febslet.2008.06.040. http://doi.wiley.com/10.1016/j.febslet.2008.06.040.
- ↑ Cloarec, Olivier; Dumas, Marc-Emmanuel; Craig, Andrew; Barton, Richard H.; Trygg, Johan; Hudson, Jane; Blancher, Christine; Gauguier, Dominique et al. (1 March 2005). "Statistical Total Correlation Spectroscopy: An Exploratory Approach for Latent Biomarker Identification from Metabolic 1 H NMR Data Sets" (in en). Analytical Chemistry 77 (5): 1282–1289. doi:10.1021/ac048630x. ISSN 0003-2700. https://pubs.acs.org/doi/10.1021/ac048630x.
- ↑ Choi, Young Hae; Kim, Hye Kyong; Hazekamp, Arno; Erkelens, Cornelis; Lefeber, Alfons W. M.; Verpoorte, Robert (1 June 2004). "Metabolomic Differentiation of Cannabis s ativa Cultivars Using 1 H NMR Spectroscopy and Principal Component Analysis" (in en). Journal of Natural Products 67 (6): 953–957. doi:10.1021/np049919c. ISSN 0163-3864. https://pubs.acs.org/doi/10.1021/np049919c.
- ↑ Wishart, D. S.; Knox, C.; Guo, A. C.; Eisner, R.; Young, N.; Gautam, B.; Hau, D. D.; Psychogios, N. et al. (1 January 2009). "HMDB: a knowledgebase for the human metabolome" (in en). Nucleic Acids Research 37 (Database): D603–D610. doi:10.1093/nar/gkn810. ISSN 0305-1048. PMC PMC2686599. PMID 18953024. https://academic.oup.com/nar/article-lookup/doi/10.1093/nar/gkn810.
- ↑ Barclay, Thomas; Ginic-Markovic, Milena; Johnston, Martin R.; Cooper, Peter; Petrovsky, Nikolai (1 January 2012). "Observation of the keto tautomer of d-fructose in D2O using 1H NMR spectroscopy" (in en). Carbohydrate Research 347 (1): 136–141. doi:10.1016/j.carres.2011.11.003. PMC PMC3254704. PMID 22129837. https://linkinghub.elsevier.com/retrieve/pii/S0008621511005404.
- ↑ 40.0 40.1 40.2 de Falco, Bruna; Incerti, Guido; Pepe, Rosa; Amato, Mariana; Lanzotti, Virginia (1 September 2016). "Metabolomic Fingerprinting of Romaneschi Globe Artichokes by NMR Spectroscopy and Multivariate Data Analysis: Metabolomics of Artichokes by NMR and Chemometrics" (in en). Phytochemical Analysis 27 (5): 304–314. doi:10.1002/pca.2632. https://onlinelibrary.wiley.com/doi/10.1002/pca.2632.
- ↑ 41.0 41.1 Wishart, David S; Guo, AnChi; Oler, Eponine; Wang, Fei; Anjum, Afia; Peters, Harrison; Dizon, Raynard; Sayeeda, Zinat et al. (7 January 2022). "HMDB 5.0: the Human Metabolome Database for 2022" (in en). Nucleic Acids Research 50 (D1): D622–D631. doi:10.1093/nar/gkab1062. ISSN 0305-1048. PMC PMC8728138. PMID 34986597. https://academic.oup.com/nar/article/50/D1/D622/6431815.
- ↑ De Almeida, Mauro V.; Couri, Mara Rubia C.; De Assis, João Vitor; Anconi, Cleber P. A.; Dos Santos, Hélio F.; De Almeida, Wagner B. (1 September 2012). "1H NMR analysis of O-methyl-inositol isomers: a joint experimental and theoretical study: A combined experimental/DFT NMR spectroscopic investigation is a powerful tool in structural/conformational analysis studies of organic compounds and" (in en). Magnetic Resonance in Chemistry 50 (9): 608–614. doi:10.1002/mrc.3848. https://onlinelibrary.wiley.com/doi/10.1002/mrc.3848.
- ↑ Stoianova-Ivanova, B.; Hadjieva, P.; Tamas̀, J. (1 August 1974). "Nonacosane-5,8-diol: A new component of plant waxes" (in en). Phytochemistry 13 (8): 1523–1525. doi:10.1016/0031-9422(74)80320-1. https://linkinghub.elsevier.com/retrieve/pii/0031942274803201.
- ↑ Lowe, Henry; Toyang, Ngeh; Steele, Blair; Valentine, Henkel; Grant, Justin; Ali, Amza; Ngwa, Wilfred; Gordon, Lorenzo (15 May 2021). "The Therapeutic Potential of Psilocybin" (in en). Molecules 26 (10): 2948. doi:10.3390/molecules26102948. ISSN 1420-3049. PMC PMC8156539. PMID 34063505. https://www.mdpi.com/1420-3049/26/10/2948.
- ↑ Adams, Roger; Pease, D. C.; Clark, J. H. (1 August 1940). "Isolation of Cannabinol, Cannabidiol and Quebrachitol from Red Oil of Minnesota Wild Hemp" (in en). Journal of the American Chemical Society 62 (8): 2194–2196. doi:10.1021/ja01865a080. ISSN 0002-7863. https://pubs.acs.org/doi/abs/10.1021/ja01865a080.
- ↑ Groce, John W.; Jones, Louis A. (1 March 1973). "Carbohydrate and cyclitol content of Cannabis" (in en). Journal of Agricultural and Food Chemistry 21 (2): 211–214. doi:10.1021/jf60186a003. ISSN 0021-8561. https://pubs.acs.org/doi/abs/10.1021/jf60186a003.
- ↑ Kortesniemi, Maaria; Sinkkonen, Jari; Yang, Baoru; Kallio, Heikki (1 March 2017). "NMR metabolomics demonstrates phenotypic plasticity of sea buckthorn (Hippophaë rhamnoides) berries with respect to growth conditions in Finland and Canada" (in en). Food Chemistry 219: 139–147. doi:10.1016/j.foodchem.2016.09.125. https://linkinghub.elsevier.com/retrieve/pii/S0308814616315205.
- ↑ 48.0 48.1 Rhodes, D; Hanson, A D (1 June 1993). "Quaternary Ammonium and Tertiary Sulfonium Compounds in Higher Plants" (in en). Annual Review of Plant Physiology and Plant Molecular Biology 44 (1): 357–384. doi:10.1146/annurev.pp.44.060193.002041. ISSN 1040-2519. https://www.annualreviews.org/doi/10.1146/annurev.pp.44.060193.002041.
- ↑ Sarma, Nandakumara D.; Waye, Andrew; ElSohly, Mahmoud A.; Brown, Paula N.; Elzinga, Sytze; Johnson, Holly E.; Marles, Robin J.; Melanson, Jeremy E. et al. (24 April 2020). "Cannabis Inflorescence for Medical Purposes: USP Considerations for Quality Attributes" (in en). Journal of Natural Products 83 (4): 1334–1351. doi:10.1021/acs.jnatprod.9b01200. ISSN 0163-3864. https://pubs.acs.org/doi/10.1021/acs.jnatprod.9b01200.
- ↑ Moyna, Patrick; Heinzen, Horacio (1 August 2001). "Improving self-defense in plants. Martial arts for vegetables" (in en). Pure and Applied Chemistry 73 (8): 1325–1330. doi:10.1351/pac200173081325. ISSN 1365-3075. https://www.degruyter.com/document/doi/10.1351/pac200173081325/html.
- ↑ Arya, Gulab Chand; Sarkar, Sutanni; Manasherova, Ekaterina; Aharoni, Asaph; Cohen, Hagai (25 June 2021). "The Plant Cuticle: An Ancient Guardian Barrier Set Against Long-Standing Rivals". Frontiers in Plant Science 12: 663165. doi:10.3389/fpls.2021.663165. ISSN 1664-462X. PMC PMC8267416. PMID 34249035. https://www.frontiersin.org/articles/10.3389/fpls.2021.663165/full.
- ↑ Inada, Noriko; Savory, Elizabeth A. (1 September 2011). "Inhibition of prepenetration processes of the powdery mildew Golovinomyces orontii on host inflorescence stems is reduced in the Arabidopsis cuticular mutant cer3 but not in cer1" (in en). Journal of General Plant Pathology 77 (5): 273–281. doi:10.1007/s10327-011-0331-0. ISSN 1345-2630. http://link.springer.com/10.1007/s10327-011-0331-0.
- ↑ Murray, A.J.S.; Ayres, P.G. (1 September 1986). "Infection with powdery mildew can enhance the accumulation of proline and glycinebetaine by salt stressed barley seedlings" (in en). Physiological and Molecular Plant Pathology 29 (2): 271–277. doi:10.1016/S0048-4059(86)80027-3. https://linkinghub.elsevier.com/retrieve/pii/S0048405986800273.
- ↑ Aslanpour, M.; Baneh, H.D.; Tehranifar, A.; Shoor, M. (2016). "The effect of Mycorrhizal fungi on the amount of glycine betaine, soluble sugar, proline, leaf water content and leaf chlorophyll of the white seedless grape under drought stress conditions". International Journal of Advanced Biotechnology and Research 7 (3): 1119-1133. https://profdoc.um.ac.ir/paper-abstract-1061692.html.
Notes
This presentation is faithful to the original, with only a few minor changes to presentation. Some grammar and punctuation was cleaned up to improve readability. In some cases important information was missing from the references, and that information was added. The original article lists references alphabetically, but this version—by design—lists them in order of appearance.